+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Procapsid of bacteriophage JBD30 computed in I4 symmetry | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | bacteriophage / virion / procapsid / VIRUS | |||||||||
| Function / homology | Bacteriophage Mu, GpT / Mu-like prophage major head subunit gpT / Bacteriophage Mu GpT domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |  Pseudomonas phage JBD30 (virus) Pseudomonas phage JBD30 (virus) | |||||||||
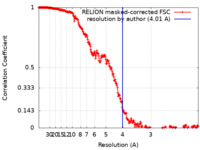
| Method | single particle reconstruction / cryo EM / Resolution: 4.01 Å | |||||||||
 Authors Authors | Valentova L / Fuzik T / Plevka P | |||||||||
| Funding support | European Union,  Czech Republic, 2 items Czech Republic, 2 items
| |||||||||
 Citation Citation |  Journal: Embo J. / Year: 2024 Journal: Embo J. / Year: 2024Title: Structure and replication of Pseudomonas aeruginosa phage JBD30 Authors: Valentova L / Plevka P / Fuzik T / Novacek J / Pospisil J | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19285.map.gz emd_19285.map.gz | 1.3 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19285-v30.xml emd-19285-v30.xml emd-19285.xml emd-19285.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19285_fsc.xml emd_19285_fsc.xml | 25.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_19285.png emd_19285.png | 257 KB | ||
| Masks |  emd_19285_msk_1.map emd_19285_msk_1.map | 1.4 GB |  Mask map Mask map | |
| Filedesc metadata |  emd-19285.cif.gz emd-19285.cif.gz | 6.1 KB | ||
| Others |  emd_19285_half_map_1.map.gz emd_19285_half_map_1.map.gz emd_19285_half_map_2.map.gz emd_19285_half_map_2.map.gz | 1.1 GB 1.1 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19285 http://ftp.pdbj.org/pub/emdb/structures/EMD-19285 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19285 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19285 | HTTPS FTP |
-Validation report
| Summary document |  emd_19285_validation.pdf.gz emd_19285_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19285_full_validation.pdf.gz emd_19285_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_19285_validation.xml.gz emd_19285_validation.xml.gz | 33.4 KB | Display | |
| Data in CIF |  emd_19285_validation.cif.gz emd_19285_validation.cif.gz | 45.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19285 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19285 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19285 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19285 | HTTPS FTP |
-Related structure data
| Related structure data |  8rkxMC  8rk3C  8rk4C  8rk5C  8rk6C  8rk7C  8rk8C  8rk9C  8rkaC  8rkbC  8rkcC  8rknC  8rkoC  8rqeC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19285.map.gz / Format: CCP4 / Size: 1.4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19285.map.gz / Format: CCP4 / Size: 1.4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.057 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19285_msk_1.map emd_19285_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
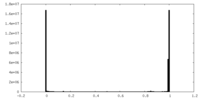
| Density Histograms |
-Half map: #1
| File | emd_19285_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
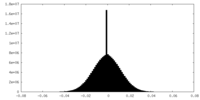
| Density Histograms |
-Half map: #2
| File | emd_19285_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
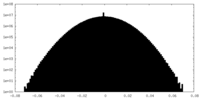
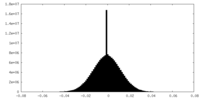
| Density Histograms |
- Sample components
Sample components
-Entire : Pseudomonas phage JBD30
| Entire | Name:  Pseudomonas phage JBD30 (virus) Pseudomonas phage JBD30 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Pseudomonas phage JBD30
| Supramolecule | Name: Pseudomonas phage JBD30 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: Phage JBD30 was propagated in P. aeruginosa strain BAA-28 and purified using CsCl gradient. NCBI-ID: 1223260 / Sci species name: Pseudomonas phage JBD30 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Molecular weight | Theoretical: 13.971 MDa |
| Virus shell | Shell ID: 1 / Name: JBD30 procapsid / Diameter: 570.0 Å / T number (triangulation number): 7 |
-Macromolecule #1: Bacteriophage Mu GpT domain-containing protein
| Macromolecule | Name: Bacteriophage Mu GpT domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage JBD30 (virus) Pseudomonas phage JBD30 (virus) |
| Molecular weight | Theoretical: 33.492707 KDa |
| Sequence | String: IITPALISAL KTSFQKHFQD ALATAPSTYL QVATVIPSTT ASNTYGWLGQ FPKLREWIGQ RVIKDMAAQG YQITNKLFES TVGVKRTDI EDDNLGVYGP LMQEMGRAAG AHPDELVFAL LKAGNANLCY DGQNFFDTDH PVYPNVDGTG TATTVSNLFA P AADPGAAW ...String: IITPALISAL KTSFQKHFQD ALATAPSTYL QVATVIPSTT ASNTYGWLGQ FPKLREWIGQ RVIKDMAAQG YQITNKLFES TVGVKRTDI EDDNLGVYGP LMQEMGRAAG AHPDELVFAL LKAGNANLCY DGQNFFDTDH PVYPNVDGTG TATTVSNLFA P AADPGAAW YLLDTSRSLK PLIYQERMKP SFTSMTKEDD EQVFMADEYR YGVRSRCNVG FGFWQLAAMS TEELNQVNFE KV YDAMRNQ KADGGRPLDI RPNLLVVPTT LRSKAKEVVG VQRLANGADN PNFELVQVLD TAWLN UniProtKB: Bacteriophage Mu GpT domain-containing protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
Details: 10 mM MgSO4, 10 mM NaCl, 50 mM Tris pH 8 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: OTHER / Details: Gatan Solarus II | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: blotting force 0, blotting time 2 s, waiting time 15 s. | ||||||||||||
| Details | phage titer 10^11 PFU |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 1-40 / Number grids imaged: 1 / Number real images: 13769 / Average exposure time: 7.5 sec. / Average electron dose: 53.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | de novo model building |
|---|---|
| Refinement | Space: REAL / Protocol: OTHER |
| Output model |  PDB-8rkx: |
 Movie
Movie Controller
Controller
























 Z
Z Y
Y X
X