[English] 日本語
 Yorodumi
Yorodumi- EMDB-18999: Cryo-EM structure of coagulation factor beta-XIIa in complex with... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of coagulation factor beta-XIIa in complex with the garadacimab Fab fragment (symmetric dimer) | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / coagulation / trypsin-like serine protease / hereditary angioedema (HAE) / BLOOD CLOTTING | |||||||||
| Function / homology |  Function and homology information Function and homology informationcoagulation factor XIIa / plasma kallikrein-kinin cascade / Factor XII activation / Defective SERPING1 causes hereditary angioedema / response to misfolded protein / positive regulation of plasminogen activation / blood coagulation, intrinsic pathway / misfolded protein binding / positive regulation of fibrinolysis / zymogen activation ...coagulation factor XIIa / plasma kallikrein-kinin cascade / Factor XII activation / Defective SERPING1 causes hereditary angioedema / response to misfolded protein / positive regulation of plasminogen activation / blood coagulation, intrinsic pathway / misfolded protein binding / positive regulation of fibrinolysis / zymogen activation / Defective factor XII causes hereditary angioedema / protein autoprocessing / positive regulation of blood coagulation / rough endoplasmic reticulum / fibrinolysis / Intrinsic Pathway of Fibrin Clot Formation / protein processing / blood coagulation / collagen-containing extracellular matrix / innate immune response / serine-type endopeptidase activity / calcium ion binding / extracellular space / extracellular exosome / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
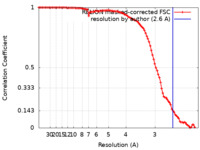
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Drulyte I | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2024 Journal: Structure / Year: 2024Title: Structural basis for the inhibition of βFXIIa by garadacimab. Authors: Ieva Drulyte / Rajesh Ghai / Saw Yen Ow / Eugene A Kapp / Adam J Quek / Con Panousis / Michael J Wilson / Andrew D Nash / Matthias Pelzing /   Abstract: Activated FXII (FXIIa) is the principal initiator of the plasma contact system and can activate both procoagulant and proinflammatory pathways. Its activity is important in the pathophysiology of ...Activated FXII (FXIIa) is the principal initiator of the plasma contact system and can activate both procoagulant and proinflammatory pathways. Its activity is important in the pathophysiology of hereditary angioedema (HAE). Here, we describe a high-resolution cryoelectron microscopy (cryo-EM) structure of the beta-chain from FXIIa (βFXIIa) complexed with the Fab fragment of garadacimab. Garadacimab binds to βFXIIa through an unusually long CDR-H3 that inserts into the S1 pocket in a non-canonical way. This structural mechanism is likely the primary contributor to the inhibition of activated FXIIa proteolytic activity in HAE. Garadacimab Fab-βFXIIa structure also reveals critical determinants of high-affinity binding of garadacimab to activated FXIIa. Structural analysis with other bona fide FXIIa inhibitors, such as benzamidine and C1-INH, reveals a surprisingly similar mechanism of βFXIIa inhibition by garadacimab. In summary, the garadacimab Fab-βFXIIa structure provides crucial insights into its mechanism of action and delineates primary and auxiliary paratopes/epitopes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18999.map.gz emd_18999.map.gz | 79 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18999-v30.xml emd-18999-v30.xml emd-18999.xml emd-18999.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18999_fsc.xml emd_18999_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_18999.png emd_18999.png | 91.2 KB | ||
| Masks |  emd_18999_msk_1.map emd_18999_msk_1.map emd_18999_msk_2.map emd_18999_msk_2.map | 83.7 MB 83.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18999.cif.gz emd-18999.cif.gz | 7 KB | ||
| Others |  emd_18999_additional_1.map.gz emd_18999_additional_1.map.gz emd_18999_additional_2.map.gz emd_18999_additional_2.map.gz emd_18999_half_map_1.map.gz emd_18999_half_map_1.map.gz emd_18999_half_map_2.map.gz emd_18999_half_map_2.map.gz | 42.4 MB 1.9 MB 77.5 MB 77.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18999 http://ftp.pdbj.org/pub/emdb/structures/EMD-18999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18999 | HTTPS FTP |
-Validation report
| Summary document |  emd_18999_validation.pdf.gz emd_18999_validation.pdf.gz | 847 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18999_full_validation.pdf.gz emd_18999_full_validation.pdf.gz | 846.6 KB | Display | |
| Data in XML |  emd_18999_validation.xml.gz emd_18999_validation.xml.gz | 17.1 KB | Display | |
| Data in CIF |  emd_18999_validation.cif.gz emd_18999_validation.cif.gz | 22.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18999 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18999 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18999 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18999 | HTTPS FTP |
-Related structure data
| Related structure data |  8r8dMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18999.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18999.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.095 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18999_msk_1.map emd_18999_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
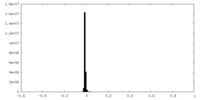
| Density Histograms |
-Mask #2
| File |  emd_18999_msk_2.map emd_18999_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
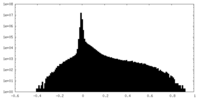
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_18999_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
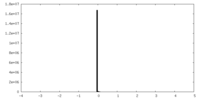
| Density Histograms |
-Additional map: Local resolution-filtered map
| File | emd_18999_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution-filtered map | ||||||||||||
| Projections & Slices |
| ||||||||||||
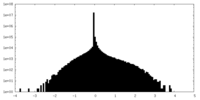
| Density Histograms |
-Half map: Half map A
| File | emd_18999_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_18999_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of coagulation factor beta-XIIa with garadacimab ...
| Entire | Name: Ternary complex of coagulation factor beta-XIIa with garadacimab Fab fragment and anti-LC-lambda VHH |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of coagulation factor beta-XIIa with garadacimab ...
| Supramolecule | Name: Ternary complex of coagulation factor beta-XIIa with garadacimab Fab fragment and anti-LC-lambda VHH type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 174 KDa |
-Macromolecule #1: Coagulation factor XII
| Macromolecule | Name: Coagulation factor XII / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.53775 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VVGGLVALRG AHPYIAALYW GHSFCAGSLI APCWVLTAAH CLQDRPAPED LTVVLGQERR NHSCEPCQTL AVRSYRLHEA FSPVSYQHD LALLRLQEDA DGSCALLSPY VQPVSLPSGA ARPSETTLCQ VAGWGHQFEG AEEYASFLQE AQVPFLSLER C SAPDVHGS ...String: VVGGLVALRG AHPYIAALYW GHSFCAGSLI APCWVLTAAH CLQDRPAPED LTVVLGQERR NHSCEPCQTL AVRSYRLHEA FSPVSYQHD LALLRLQEDA DGSCALLSPY VQPVSLPSGA ARPSETTLCQ VAGWGHQFEG AEEYASFLQE AQVPFLSLER C SAPDVHGS SILPGMLCAG FLEGGTDACQ GDSGGPLVCE DQAAERRLTL QGIISWGSGC GDRNKPGVYT DVAYYLAWIR EH TVSEPEA UniProtKB: Coagulation factor XII |
-Macromolecule #2: Garadacimab heavy chain variable region
| Macromolecule | Name: Garadacimab heavy chain variable region / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.120281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLLESGGG LVQPGGSLRL SCAASGFTFS KYIMQWVRQA PGKGLEWVSG IDIPTKGTVY ADSVKGRFTI SRDNSKNTLY LQMNSLRAE DTAVYYCARA LPRSGYLISP HYYYYALDVW GQGTTVTVSS ASTKGPSVFP LAPCSRSTSE STAALGCLVK D YFPEPVTV ...String: EVQLLESGGG LVQPGGSLRL SCAASGFTFS KYIMQWVRQA PGKGLEWVSG IDIPTKGTVY ADSVKGRFTI SRDNSKNTLY LQMNSLRAE DTAVYYCARA LPRSGYLISP HYYYYALDVW GQGTTVTVSS ASTKGPSVFP LAPCSRSTSE STAALGCLVK D YFPEPVTV SWNSGALTSG VHTFPAVLQS SGLYSLSSVV TVPSSSLGTK TYTCNVDHKP SNTKVDKRVE SKYGPP |
-Macromolecule #3: Garadacimab light chain variable region
| Macromolecule | Name: Garadacimab light chain variable region / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.762174 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSVLTQPPSA SGTPGQRVTI SCSGSSSNIG RNYVYWYQQL PGTAPKLLIY SNNQRPSGVP DRFSGSKSGT SASLAISGLR SEDEADYYC AAWDASLRGV FGGGTKLTVL GQPKAAPSVT LFPPSSEELQ ANKATLVCLI SDFYPGAVTV AWKADSSPVK A GVETTTPS ...String: QSVLTQPPSA SGTPGQRVTI SCSGSSSNIG RNYVYWYQQL PGTAPKLLIY SNNQRPSGVP DRFSGSKSGT SASLAISGLR SEDEADYYC AAWDASLRGV FGGGTKLTVL GQPKAAPSVT LFPPSSEELQ ANKATLVCLI SDFYPGAVTV AWKADSSPVK A GVETTTPS KQSNNKYAAS SYLSLTPEQW KSHRSYSCQV THEGSTVEKT VAPTECS |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 18 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 5.5 / Details: 10 mM Na Acetate, 100 mM NaCl pH 5.5 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: plasma current 20 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | Immediately before blotting and plunge freezing, fluorinated octyl maltoside (FOM) was added to the sample to the final concentration of 0.005%-0.01% (w/v) |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris / Energy filter - Slit width: 10 eV Details: Electron source E-CFEG (cold-FEG), energy filter Selectris X |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 2 / Number real images: 4000 / Average electron dose: 50.0 e/Å2 / Details: 2000 with 0.005% FOM and 2000 with 0.01% FOM |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.25 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)