[English] 日本語
 Yorodumi
Yorodumi- EMDB-18407: Cryo-EM structure of SidH from Legionella pneumophila in complex ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SidH from Legionella pneumophila in complex with LubX | |||||||||
 Map data Map data | EM Map of Legionella effector SidH bound to metaeffector LubX | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SidH / Legionella / tRNA-binding / EF-Tu-binding / RNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationguanosine tetraphosphate binding / translation elongation factor activity / ubiquitin-protein transferase activity / protein ubiquitination / GTPase activity / GTP binding / cytosol Similarity search - Function | |||||||||
| Biological species |   | |||||||||
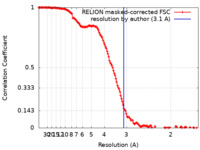
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Sharma R / Adams M / Bhogaraju S | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural basis for the toxicity of Legionella pneumophila effector SidH. Authors: Rahul Sharma / Michael Adams / Simonne Griffith-Jones / Tobias Sahr / Laura Gomez-Valero / Felix Weis / Michael Hons / Sarah Gharbi / Rayene Berkane / Alexandra Stolz / Carmen Buchrieser / Sagar Bhogaraju /   Abstract: Legionella pneumophila (LP) secretes more than 300 effectors into the host cytosol to facilitate intracellular replication. One of these effectors, SidH, 253 kDa in size with no sequence similarity ...Legionella pneumophila (LP) secretes more than 300 effectors into the host cytosol to facilitate intracellular replication. One of these effectors, SidH, 253 kDa in size with no sequence similarity to proteins of known function is toxic when overexpressed in host cells. SidH is regulated by the LP metaeffector LubX which targets SidH for degradation in a temporal manner during LP infection. The mechanism underlying the toxicity of SidH and its role in LP infection are unknown. Here, we determined the cryo-EM structure of SidH at 2.7 Å revealing a unique alpha helical arrangement with no overall similarity to known protein structures. Surprisingly, purified SidH came bound to a E. coli EF-Tu/t-RNA/GTP ternary complex which could be modeled into the cryo-EM density. Mutation of residues disrupting the SidH-tRNA interface and SidH-EF-Tu interface abolish the toxicity of overexpressed SidH in human cells, a phenotype confirmed in infection of Acanthamoeba castellani. We also present the cryo-EM structure of SidH in complex with a U-box domain containing ubiquitin ligase LubX delineating the mechanism of regulation of SidH. Our data provide the basis for the toxicity of SidH and into its regulation by the metaeffector LubX. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18407.map.gz emd_18407.map.gz | 149.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18407-v30.xml emd-18407-v30.xml emd-18407.xml emd-18407.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18407_fsc.xml emd_18407_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_18407.png emd_18407.png | 36.3 KB | ||
| Filedesc metadata |  emd-18407.cif.gz emd-18407.cif.gz | 8.1 KB | ||
| Others |  emd_18407_half_map_1.map.gz emd_18407_half_map_1.map.gz emd_18407_half_map_2.map.gz emd_18407_half_map_2.map.gz | 194 MB 194.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18407 http://ftp.pdbj.org/pub/emdb/structures/EMD-18407 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18407 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18407 | HTTPS FTP |
-Validation report
| Summary document |  emd_18407_validation.pdf.gz emd_18407_validation.pdf.gz | 675.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18407_full_validation.pdf.gz emd_18407_full_validation.pdf.gz | 675.1 KB | Display | |
| Data in XML |  emd_18407_validation.xml.gz emd_18407_validation.xml.gz | 21.5 KB | Display | |
| Data in CIF |  emd_18407_validation.cif.gz emd_18407_validation.cif.gz | 27.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18407 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18407 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18407 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18407 | HTTPS FTP |
-Related structure data
| Related structure data |  8qhcMC  8qfsC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18407.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18407.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM Map of Legionella effector SidH bound to metaeffector LubX | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.827 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map 1
| File | emd_18407_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_18407_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Quaternary complex of SidH with E.coli tRNA, EF-Tu and LubX
| Entire | Name: Quaternary complex of SidH with E.coli tRNA, EF-Tu and LubX |
|---|---|
| Components |
|
-Supramolecule #1: Quaternary complex of SidH with E.coli tRNA, EF-Tu and LubX
| Supramolecule | Name: Quaternary complex of SidH with E.coli tRNA, EF-Tu and LubX type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Elongation factor Tu
| Macromolecule | Name: Elongation factor Tu / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.370492 KDa |
| Sequence | String: MSKEKFERTK PHVNVGTIGH VDHGKTTLTA AITTVLAKTY GGAARAFDQI DNAPEEKARG ITINTSHVEY DTPTRHYAHV DCPGHADYV KNMITGAAQM DGAILVVAAT DGPMPQTREH ILLGRQVGVP YIIVFLNKCD MVDDEELLEL VEMEVRELLS Q YDFPGDDT ...String: MSKEKFERTK PHVNVGTIGH VDHGKTTLTA AITTVLAKTY GGAARAFDQI DNAPEEKARG ITINTSHVEY DTPTRHYAHV DCPGHADYV KNMITGAAQM DGAILVVAAT DGPMPQTREH ILLGRQVGVP YIIVFLNKCD MVDDEELLEL VEMEVRELLS Q YDFPGDDT PIVRGSALKA LEGDAEWEAK ILELAGFLDS YIPEPERAID KPFLLPIEDV FSISGRGTVV TGRVERGIIK VG EEVEIVG IKETQKSTCT GVEMFRKLLD EGRAGENVGV LLRGIKREEI ERGQVLAKPG TIKPHTKFES EVYILSKDEG GRH TPFFKG YRPQFYFRTT DVTGTIELPE GVEMVMPGDN IKMVVTLIHP IAMDDGLRFA IREGGRTVGA GVVAKVLS UniProtKB: Elongation factor Tu |
-Macromolecule #3: Protein SidH
| Macromolecule | Name: Protein SidH / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 255.113719 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLVPRGSM KRTIETYIIY LKEDLKIADT CKTIKDGLLK SITDKTHFSE ELATYFERDN PNAPFKVNTT DPTQVAVLK KLLNALENAE KSFRAIENID ISRDRYTAMI AKDAVMVSYK AVHEIYAALQ LINHSNSDIQ DIVGPHIQKL L PQMALASK ...String: MGSSHHHHHH SSGLVPRGSM KRTIETYIIY LKEDLKIADT CKTIKDGLLK SITDKTHFSE ELATYFERDN PNAPFKVNTT DPTQVAVLK KLLNALENAE KSFRAIENID ISRDRYTAMI AKDAVMVSYK AVHEIYAALQ LINHSNSDIQ DIVGPHIQKL L PQMALASK ALGNFAPEHP EESAGAVLAG VVNMLPTEKP TESESLGKLS NLIFELPHYF EELQKLIATG ASGIATKSIT SA EDYQSAM IKKANETKYY FEQLSSKSGL LAIPSYLSIV KRLIAHSTDL VNAGAPLTKQ AYLDAVAKLE DIKHNILPQL ISE LEMVEE SMGLKPGLLT DPALEQMNKY YTQLAEQVDN IAKAAGVLDT VSDYSDSIGG KIVRFLAGDS KKLDVGPKLT PAPD LGVLM DDVFIQKRRS NQESRLNEAR LSSEDKSVLA AANRFFDKIG SYNSIHKAWS KWSLANISQS EKDALIKEYK QFQPH FAAL YPDIDKLVVD ALTQPTGSNI VSRLYSSDYK QLWSSDHFKQ VLSCKDSVLS SIQQSLAQSE FKAKLIEKTM SHSEET AYS MNNKTTNLTT RVQPFEPLKF TLEDDKPVEY YHKRVIAASN QILELERAQK GVAEFFNYIQ KKYPHENPSF DSLDESD KE FLRKAYKTFQ PQLLALKHDD INTRLVSSLT SSKPTDPPLR LTDLVSLKSG INDYLNEKIS DLNQDKTTLL DKEEEARE E QYAKNPLVAK GAELEKQTLF GQMSKLKLSK SVDDFFNKKF QTYLKDNLSP EVWKQLSSNG ETLDFDKIPY LEFHKDSPE VAMYKQLINS MHYMKNGLEK LESLNDYGDP NNIYHRTRFV MTTFNALVMN ICFSKYYVME AGNNPGLKAI VQEGLDLLKP LEGMPLIGD YLKTTEKQEP PKQNIITAWK KQQAVVESGL SKGPKPKTDQ QLISEQLGKI QEAIDNFDGD LEVSDSAREK I KTQIGEFA KGISGLSFGP GSVKKILAAL TKLETQLSNL DKESPEVTLG KLKDIHSELN AQFRAAAEYT EYHSGQKFGS YS NNISTIV SNFCNGLVSN LPLKQEPAPK VKAPEKPVTP VITGTTNPHE VVFGTKHEEF NSVYQPYLLL KRITDEFRDQ NNP YKPSFD ELKEEAVSYY DKIQPLLESV DPKFDKNFIA KHKESSTLLK AIDEVMSMRQ RINNPESSFA KLKDLHLEGD FEKE ENKEK FRQLYKEIQP YLHKIDSTYD QTQFLEGLKT AKDFSGALQR IMNEENALQQ STSLKDTSYL QLIAESLYQI PVKLN KLKA EPDTPEPSKE EIDANVKAFV EGLNGLSFGP GSVKKILSTA AKLQMQLSDI GKEGRELTMG RLKEIQAEFG TILMAA ADN AEFHLGLKPG TYSRTVSERF EKFYSSLIVN LPLEKDQTGL ELLIDTTSTQ KRLAREMERL ESVKEDTSAI DTKKSIF GT EHEQFSTLYQ PYASLRHIAK DIEDGVHMNL YERTLEELKE EASNEYKKIQ PYLAKINPEF TEDYISKTDG EYSLLHAI D RVFEERHKIN KPSSPFDKLR DLYLDGDFEK EENKEQFLQL YAELQPHLIK INYQYDLAYF LRELQTPEDF KAATERIIN DESKLQELIT GLDDTKRLKV KLCEERIGYF IDLLKKQELE VGPEKIQAFK EKIFFNYIHA NVNSALDAKI GSHAEQFLQF IEKDFLDKK NEILEKITID QDMEEEIAKA IDRIAPDIIN NKIESFKKLL FDSYIQSDIK NSLHNELGIY TSLFIDKISP E IHLHQSEI LGNVAFDSKM GAEIGSKINA ITPGILLNNN PLKEAYVDLN NTLKEINTLL DEENKKTRDN PCRNEKIAKL TS LKDRLSD LDSIPKENTV EFLKKMQEET RSSLKSLESN DALINIYDVL NSLKETIENG SDLPEIKKDK LQMISDVQNI LSN FDKNPA ERLNLAVQIL NDSNPEVLSK TKGNFLIGEA FKGQVFTNYI NTKISEQLNN ELGPYGKVFL KQIMPDFIAK KSEI IKEIA IDNMETGLET QFKIHAPAIF EKNKELKAAY EQLNVHLKEV QSLIEAEEKK PKGNPCREEK IAALRSHQSQ LMNTQ RIPD HETLRFLQEQ NKSAKSFMGK LEKYDTMISV YDSLTEIREH VSNHKSLSKE IKDEKIQEIS KMEDMLKTTS KEPSIR LAE VKAHGLSDQC KNVLLKNSDN FLVSFFKTLF SKLFNIKNEN ETLVSSFKQR LQNIKGPEPV ATPMETPENE APLVNAN IT RF UniProtKB: SidH |
-Macromolecule #4: E3 ubiquitin--protein ligase
| Macromolecule | Name: E3 ubiquitin--protein ligase / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.50683 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHSAG LEVLFQGPMG YRIEMATRNP FDIDHKSKYL REAALEANLS HPETTPTMLT CPIDSGFLKD PVITPEGFVY NKSSILKWL ETKKEDPQSR KPLTAKDLQP FPELLIIVNR FVETQTNYEK LKNRLVQNAR VAARQKEYTE IPDIFLCPIS K TLIKTPVI ...String: MHHHHHHSAG LEVLFQGPMG YRIEMATRNP FDIDHKSKYL REAALEANLS HPETTPTMLT CPIDSGFLKD PVITPEGFVY NKSSILKWL ETKKEDPQSR KPLTAKDLQP FPELLIIVNR FVETQTNYEK LKNRLVQNAR VAARQKEYTE IPDIFLCPIS K TLIKTPVI TAQGKVYDQE ALSNFLIATG NKDETGKKLS IDDVVVFDEL YQQIKVYNFY RKREMQKNQI QPSVSSGFGF FS LNFLTSW LWGTEEKKEK TSSDMTY UniProtKB: E3 ubiquitin--protein ligase |
-Macromolecule #2: t-RNA
| Macromolecule | Name: t-RNA / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.620766 KDa |
| Sequence | String: GCCCGGA(4SU)AG CUCAG(H2U)CGG(H2U) AGAGCAGGGG AUUGAAAAUC CCCGUG(3AU)CCU UGG(5MU)(PSU)C GAU UCCGAGUCCG GGCACCA |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 1 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 43.37 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X