+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of apo Clostridioides difficile toxin B | ||||||||||||||||||||||||||||||||||||
 Map data Map data | overall map used for defining masked regions | ||||||||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||||||||
 Keywords Keywords | microbiology / pore forming region / CROP dynamics / TOXIN / Structural Genomics / Structural Genomics Consortium / SGC | ||||||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationglucosyltransferase activity / host cell cytosol / Transferases; Glycosyltransferases; Hexosyltransferases / cysteine-type peptidase activity / host cell endosome membrane / toxin activity / Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases / lipid binding / host cell plasma membrane / proteolysis ...glucosyltransferase activity / host cell cytosol / Transferases; Glycosyltransferases; Hexosyltransferases / cysteine-type peptidase activity / host cell endosome membrane / toxin activity / Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases / lipid binding / host cell plasma membrane / proteolysis / extracellular region / membrane / metal ion binding Similarity search - Function | ||||||||||||||||||||||||||||||||||||
| Biological species |  Priestia megaterium DSM 319 (bacteria) / Priestia megaterium DSM 319 (bacteria) /  Clostridioides difficile (bacteria) Clostridioides difficile (bacteria) | ||||||||||||||||||||||||||||||||||||
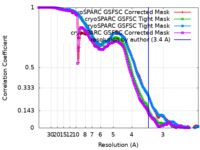
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||||||||||||||||||||||||||
 Authors Authors | Kinsolving J / Bous J / Structural Genomics Consortium (SGC) | ||||||||||||||||||||||||||||||||||||
| Funding support |  Sweden, Sweden,  Denmark, 11 items Denmark, 11 items
| ||||||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Structural and functional insight into the interaction of Clostridioides difficile toxin B and FZD. Authors: Julia Kinsolving / Julien Bous / Pawel Kozielewicz / Sara Košenina / Rawan Shekhani / Lukas Grätz / Geoffrey Masuyer / Yuankai Wang / Pål Stenmark / Min Dong / Gunnar Schulte /   Abstract: The G protein-coupled receptors of the Frizzled (FZD) family, in particular FZD, are receptors that are exploited by Clostridioides difficile toxin B (TcdB), the major virulence factor responsible ...The G protein-coupled receptors of the Frizzled (FZD) family, in particular FZD, are receptors that are exploited by Clostridioides difficile toxin B (TcdB), the major virulence factor responsible for pathogenesis associated with Clostridioides difficile infection. We employ a live-cell assay examining the affinity between full-length FZDs and TcdB. Moreover, we present cryoelectron microscopy structures of TcdB alone and in complex with full-length FZD, which reveal that large structural rearrangements of the combined repetitive polypeptide domain are required for interaction with FZDs and other TcdB receptors, constituting a first step for receptor recognition. Furthermore, we show that bezlotoxumab, an FDA-approved monoclonal antibody to treat Clostridioides difficile infection, favors the apo-TcdB structure and thus disrupts binding with FZD. The dynamic transition between the two conformations of TcdB also governs the stability of the pore-forming region. Thus, our work provides structural and functional insight into how conformational dynamics of TcdB determine receptor binding. | ||||||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18373.map.gz emd_18373.map.gz | 447.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18373-v30.xml emd-18373-v30.xml emd-18373.xml emd-18373.xml | 24.9 KB 24.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18373_fsc.xml emd_18373_fsc.xml emd_18373_fsc_2.xml emd_18373_fsc_2.xml emd_18373_fsc_3.xml emd_18373_fsc_3.xml emd_18373_fsc_4.xml emd_18373_fsc_4.xml | 16.5 KB 16.5 KB 16.5 KB 16.5 KB | Display Display Display Display |  FSC data file FSC data file |
| Images |  emd_18373.png emd_18373.png | 77.1 KB | ||
| Masks |  emd_18373_msk_1.map emd_18373_msk_1.map emd_18373_msk_2.map emd_18373_msk_2.map | 476.8 MB 476.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18373.cif.gz emd-18373.cif.gz | 8.2 KB | ||
| Others |  emd_18373_half_map_1.map.gz emd_18373_half_map_1.map.gz emd_18373_half_map_2.map.gz emd_18373_half_map_2.map.gz | 441.9 MB 441.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18373 http://ftp.pdbj.org/pub/emdb/structures/EMD-18373 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18373 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18373 | HTTPS FTP |
-Validation report
| Summary document |  emd_18373_validation.pdf.gz emd_18373_validation.pdf.gz | 1.3 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18373_full_validation.pdf.gz emd_18373_full_validation.pdf.gz | 1.3 MB | Display | |
| Data in XML |  emd_18373_validation.xml.gz emd_18373_validation.xml.gz | 25.8 KB | Display | |
| Data in CIF |  emd_18373_validation.cif.gz emd_18373_validation.cif.gz | 34.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18373 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18373 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18373 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18373 | HTTPS FTP |
-Related structure data
| Related structure data |  8qenMC  8qeoC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18373.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18373.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | overall map used for defining masked regions | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1592 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18373_msk_1.map emd_18373_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
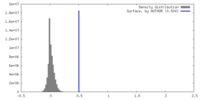
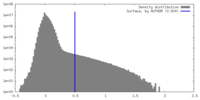
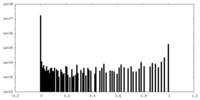
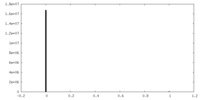
| Density Histograms |
-Mask #2
| File |  emd_18373_msk_2.map emd_18373_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18373_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18373_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : apo-structure of Clostridioides difficile toxin B
| Entire | Name: apo-structure of Clostridioides difficile toxin B |
|---|---|
| Components |
|
-Supramolecule #1: apo-structure of Clostridioides difficile toxin B
| Supramolecule | Name: apo-structure of Clostridioides difficile toxin B / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Priestia megaterium DSM 319 (bacteria) Priestia megaterium DSM 319 (bacteria) |
-Supramolecule #2: Masked region 1
| Supramolecule | Name: Masked region 1 / type: organelle_or_cellular_component / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Priestia megaterium DSM 319 (bacteria) Priestia megaterium DSM 319 (bacteria) |
-Supramolecule #3: Masked region 2
| Supramolecule | Name: Masked region 2 / type: organelle_or_cellular_component / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Priestia megaterium DSM 319 (bacteria) Priestia megaterium DSM 319 (bacteria) |
-Macromolecule #1: Toxin B
| Macromolecule | Name: Toxin B / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Clostridioides difficile (bacteria) Clostridioides difficile (bacteria) |
| Molecular weight | Theoretical: 273.551562 KDa |
| Recombinant expression | Organism:  Priestia megaterium DSM 319 (bacteria) Priestia megaterium DSM 319 (bacteria) |
| Sequence | String: MDKLVHLNQR GKCTMSLVNR KQLEKMANVR FRTQEDEYVA ILDALEEYHN MSENTVVEKY LKLKDINSLT DIYIDTYKKS GRNKALKKF KEYLVTEVLE LKNNNLTPVE KNLHFVWIGG QINDTAINYI NQWKDVNSDY NVNVFYDSNA FLINTLKKTV V ESAINDTL ...String: MDKLVHLNQR GKCTMSLVNR KQLEKMANVR FRTQEDEYVA ILDALEEYHN MSENTVVEKY LKLKDINSLT DIYIDTYKKS GRNKALKKF KEYLVTEVLE LKNNNLTPVE KNLHFVWIGG QINDTAINYI NQWKDVNSDY NVNVFYDSNA FLINTLKKTV V ESAINDTL ESFRENLNDP RFDYNKFFRK RMEIIYDKQK NFINYYKAQR EENPELIIDD IVKTYLSNEY SKEIDELNTY IE ESLNKIT QNSGNDVRNF EEFKNGESFN LYEQELVERW NLAAASDILR ISALKEIGGM YLDVDMLPGI QPDLFESIEK PSS VTVDFW EMTKLEAIMK YKEYIPEYTS EHFDMLDEEV QSSFESVLAS KSDKSEIFSS LGDMEASPLE VKIAFNSKGI INQG LISVK DSYCSNLIVK QIENRYKILN NSLNPAISED NDFNTTTNTF IDSIMAEANA DNGRFMMELG KYLRVGFFPD VKTTI NLSG PEAYAAAYQD LLMFKEGSMN IHLIEADLRN FEISKTNISQ STEQEMASLW SFDDARAKAQ FEEYKRNYFE GSLGED DNL DFSQNIVVDK EYLLEKISSL ARSSERGYIH YIVQLQGDKI SYEAACNLFA KTPYDSVLFQ KNIEDSEIAY YYNPGDG EI QEIDKYKIPS IISDRPKIKL TFIGHGKDEF NTDIFAGFDV DSLSTEIEAA IDLAKEDISP KSIEINLLGC NMFSYSIN V EETYPGKLLL KVKDKISELM PSISQDSIIV SANQYEVRIN SEGRRELLDH SGEWINKEES IIKDISSKEY ISFNPKENK ITVKSKNLPE LSTLLQEIRN NSNSSDIELE EKVMLTECEI NVISNIDTQI VEERIEEAKN LTSDSINYIK DEFKLIESIS DALCDLKQQ NELEDSHFIS FEDISETDEG FSIRFINKET GESIFVETEK TIFSEYANHI TEEISKIKGT IFDTVNGKLV K KVNLDTTH EVNTLNAAFF IQSLIEYNSS KESLSNLSVA MKVQVYAQLF STGLNTITDA AKVVELVSTA LDETIDLLPT LS EGLPIIA TIIDGVSLGA AIKELSETSD PLLRQEIEAK IGIMAVNLTT ATTAIITSSL GIASGFSILL VPLAGISAGI PSL VNNELV LRDKATKVVD YFKHVSLVET EGVFTLLDDK IMMPQDDLVI SEIDFNNNSI VLGKCEIWRM EGGSGHTVTD DIDH FFSAP SITYREPHLS IYDVLEVQKE ELDLSKDLMV LPNAPNRVFA WETGWTPGLR SLENDGTKLL DRIRDNYEGE FYWRY FAFI ADALITTLKP RYEDTNIRIN LDSNTRSFIV PIITTEYIRE KLSYSFYGSG GTYALSLSQY NMGINIELSE SDVWII DVD NVVRDVTIES DKIKKGDLIE GILSTLSIEE NKIILNSHEI NFSGEVNGSN GFVSLTFSIL EGINAIIEVD LLSKSYK LL ISGELKILML NSNHIQQKID YIGFNSELQK NIPYSFVDSE GKENGFINGS TKEGLFVSEL PDVVLISKVY MDDSKPSF G YYSNNLKDVK VITKDNVNIL TGYYLKDDIK ISLSLTLQDE KTIKLNSVHL DESGVAEILK FMNRKGNTNT SDSLMSFLE SMNIKSIFVN FLQSNIKFIL DANFIISGTT SIGQFEFICD ENDNIQPYFI KFNTLETNYT LYVGNRQNMI VEPNYDLDDS GDISSTVIN FSQKYLYGID SCVNKVVISP NIYTDEINIT PVYETNNTYP EVIVLDANYI NEKINVNIND LSIRYVWSND G NDFILMST SEENKVSQVK IRFVNVFKDK TLANKLSFNF SDKQDVPVSE IILSFTPSYY EDGLIGYDLG LVSLYNEKFY IN NFGMMVS GLIYINDSLY YFKPPVNNLI TGFVTVGDDK YYFNPINGGA ASIGETIIDD KNYYFNQSGV LQTGVFSTED GFK YFAPAN TLDENLEGEA IDFTGKLIID ENIYYFDDNY RGAVEWKELD GEMHYFSPET GKAFKGLNQI GDYKYYFNSD GVMQ KGFVS INDNKHYFDD SGVMKVGYTE IDGKHFYFAE NGEMQIGVFN TEDGFKYFAH HNEDLGNEEG EEISYSGILN FNNKI YYFD DSFTAVVGWK DLEDGSKYYF DEDTAEAYIG LSLINDGQYY FNDDGIMQVG FVTINDKVFY FSDSGIIESG VQNIDD NYF YIDDNGIVQI GVFDTSDGYK YFAPANTVND NIYGQAVEYS GLVRVGEDVY YFGETYTIET GWIYDMENES DKYYFNP ET KKACKGINLI DDIKYYFDEK GIMRTGLISF ENNNYYFNEN GEMQFGYINI EDKMFYFGED GVMQIGVFNT PDGFKYFA H QNTLDENFEG ESINYTGWLD LDEKRYYFTD EYIAATGSVI IDGEEYYFDP DTAQLVISEG YRPHAGLRGS HHHHHH UniProtKB: Toxin B |
-Macromolecule #2: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 2 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.630 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 100 mM TRIS-HCl pH 7.5 200 mM NaCl 0.002% LMNG 0.0002% CHS 0.0002% GDN | ||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.101 kPa | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 17269 / Average exposure time: 3.0 sec. / Average electron dose: 50.453 e/Å2 Details: Images collected in super-resolution mode, faster acquisition mode, with 4 exposures per hole |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 105000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-8qen: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)