[English] 日本語
 Yorodumi
Yorodumi- EMDB-18246: Outer kinetochore Dam1 protomer dimer Ndc80-Nuf2 coiled-coil complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | 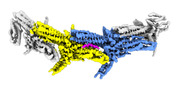 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Outer kinetochore Dam1 protomer dimer Ndc80-Nuf2 coiled-coil complex | |||||||||
 Map data Map data | Local resolution filtered map used for model refinement | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kinetochore / microtubule / error correction / chromosome segregation / CELL CYCLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationmitotic spindle polar microtubule / Ndc80 complex / DASH complex / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / kinetochore organization / positive regulation of attachment of spindle microtubules to kinetochore / meiotic chromosome segregation / mitotic spindle pole body / mitotic spindle midzone ...mitotic spindle polar microtubule / Ndc80 complex / DASH complex / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / kinetochore organization / positive regulation of attachment of spindle microtubules to kinetochore / meiotic chromosome segregation / mitotic spindle pole body / mitotic spindle midzone / attachment of spindle microtubules to kinetochore / condensed chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / microtubule plus-end binding / spindle pole body / protein localization to kinetochore / positive regulation of microtubule polymerization / mitotic spindle organization / chromosome segregation / spindle microtubule / mitotic spindle / kinetochore / spindle pole / microtubule / cell division / protein-containing complex binding / identical protein binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Muir KW / Barford D | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Structural mechanism of outer kinetochore Dam1-Ndc80 complex assembly on microtubules. Authors: Kyle W Muir / Christopher Batters / Tom Dendooven / Jing Yang / Ziguo Zhang / Alister Burt / David Barford /  Abstract: Kinetochores couple chromosomes to the mitotic spindle to segregate the genome during cell division. An error correction mechanism drives the turnover of kinetochore-microtubule attachments until ...Kinetochores couple chromosomes to the mitotic spindle to segregate the genome during cell division. An error correction mechanism drives the turnover of kinetochore-microtubule attachments until biorientation is achieved. The structural basis for how kinetochore-mediated chromosome segregation is accomplished and regulated remains an outstanding question. In this work, we describe the cryo-electron microscopy structure of the budding yeast outer kinetochore Ndc80 and Dam1 ring complexes assembled onto microtubules. Complex assembly occurs through multiple interfaces, and a staple within Dam1 aids ring assembly. Perturbation of key interfaces suppresses yeast viability. Force-rupture assays indicated that this is a consequence of impaired kinetochore-microtubule attachment. The presence of error correction phosphorylation sites at Ndc80-Dam1 ring complex interfaces and the Dam1 staple explains how kinetochore-microtubule attachments are destabilized and reset. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18246.map.gz emd_18246.map.gz | 20.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18246-v30.xml emd-18246-v30.xml emd-18246.xml emd-18246.xml | 31.2 KB 31.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_18246.png emd_18246.png | 60.8 KB | ||
| Filedesc metadata |  emd-18246.cif.gz emd-18246.cif.gz | 8 KB | ||
| Others |  emd_18246_additional_1.map.gz emd_18246_additional_1.map.gz emd_18246_half_map_1.map.gz emd_18246_half_map_1.map.gz emd_18246_half_map_2.map.gz emd_18246_half_map_2.map.gz | 119.9 MB 226.4 MB 226.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18246 http://ftp.pdbj.org/pub/emdb/structures/EMD-18246 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18246 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18246 | HTTPS FTP |
-Validation report
| Summary document |  emd_18246_validation.pdf.gz emd_18246_validation.pdf.gz | 885.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18246_full_validation.pdf.gz emd_18246_full_validation.pdf.gz | 884.7 KB | Display | |
| Data in XML |  emd_18246_validation.xml.gz emd_18246_validation.xml.gz | 16.1 KB | Display | |
| Data in CIF |  emd_18246_validation.cif.gz emd_18246_validation.cif.gz | 19.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18246 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18246 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18246 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18246 | HTTPS FTP |
-Related structure data
| Related structure data |  8q84MC  8q85C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18246.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18246.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered map used for model refinement | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened 3D reconstruction
| File | emd_18246_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened 3D reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18246_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18246_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Outer kinetochore Dam1 protomer dimer with staple and Ndc80-Nuf2 ...
+Supramolecule #1: Outer kinetochore Dam1 protomer dimer with staple and Ndc80-Nuf2 ...
+Macromolecule #1: Kinetochore protein NDC80
+Macromolecule #2: Kinetochore protein NUF2
+Macromolecule #3: DASH complex subunit DAM1
+Macromolecule #4: DASH complex subunit DUO1
+Macromolecule #5: DASH complex subunit DAD2
+Macromolecule #6: DASH complex subunit DAD1
+Macromolecule #7: DASH complex subunit DAD4
+Macromolecule #8: DASH complex subunit DAD3
+Macromolecule #9: DASH complex subunit SPC34
+Macromolecule #10: DASH complex subunit ASK1
+Macromolecule #11: DASH complex subunit HSK3
+Macromolecule #12: DASH complex subunit SPC19
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.15 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 248732 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD Software: (Name: cryoSPARC (ver. 3.3.2), RELION (ver. 4.0-dev)) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 4.2) |
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Details | Initial rigid body fitting was performed in chimera, with manual correction in coot and real-space refinement in PHENIX |
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 540.98 |
| Output model |  PDB-8q84: |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X






















 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)
