+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | 48S late-stage initiation complex with non methylated mRNA | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | translation initiation / ribosome / non methylated mRNA / TRANSLATION | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Major pathway of rRNA processing in the nucleolus and cytosol / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) ...Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Major pathway of rRNA processing in the nucleolus and cytosol / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / eukaryotic translation initiation factor 2 complex / eukaryotic 48S preinitiation complex / laminin receptor activity / positive regulation of signal transduction by p53 class mediator / phagocytic cup / ubiquitin ligase inhibitor activity / ribosomal small subunit binding / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / 90S preribosome / regulation of translational fidelity / ribosomal subunit export from nucleus / ribosomal small subunit export from nucleus / translation regulator activity / translational termination / laminin binding / cytosolic ribosome / rough endoplasmic reticulum / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / gastrulation / MDM2/MDM4 family protein binding / translation initiation factor activity / class I DNA-(apurinic or apyrimidinic site) endonuclease activity / DNA-(apurinic or apyrimidinic site) lyase / rescue of stalled ribosome / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cellular response to leukemia inhibitory factor / maturation of SSU-rRNA / small-subunit processome / translational initiation / protein kinase C binding / positive regulation of apoptotic signaling pathway / positive regulation of protein-containing complex assembly / spindle / cytoplasmic stress granule / modification-dependent protein catabolic process / rRNA processing / protein tag activity / ribosomal small subunit biogenesis / positive regulation of canonical Wnt signaling pathway / rhythmic process / small ribosomal subunit rRNA binding / ribosome binding / regulation of translation / virus receptor activity / ribosomal small subunit assembly / small ribosomal subunit / cytosolic small ribosomal subunit / perikaryon / cytosolic large ribosomal subunit / cytoplasmic translation / mitochondrial inner membrane / postsynaptic density / cell differentiation / rRNA binding / ribosome / protein ubiquitination / structural constituent of ribosome / ribonucleoprotein complex / iron ion binding / translation / positive regulation of protein phosphorylation / cell division / DNA repair / mRNA binding / centrosome / ubiquitin protein ligase binding / dendrite / synapse / nucleolus / apoptotic process / perinuclear region of cytoplasm / Golgi apparatus / ATP hydrolysis activity / DNA binding / RNA binding / zinc ion binding / ATP binding / membrane / nucleus / metal ion binding / plasma membrane / cytosol / cytoplasm 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
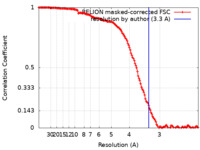
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.3 Å | |||||||||
 データ登録者 データ登録者 | Guca E / Lima LHF / Boissier F / Hashem Y | |||||||||
| 資金援助 | European Union, 1件
| |||||||||
 引用 引用 |  ジャーナル: Mol Cell / 年: 2024 ジャーナル: Mol Cell / 年: 2024タイトル: N-methyladenosine in 5' UTR does not promote translation initiation. 著者: Ewelina Guca / Rodrigo Alarcon / Michael Z Palo / Leonardo Santos / Santiago Alonso-Gil / Marcos Davyt / Leonardo H F de Lima / Fanny Boissier / Sarada Das / Bojan Zagrovic / Joseph D Puglisi ...著者: Ewelina Guca / Rodrigo Alarcon / Michael Z Palo / Leonardo Santos / Santiago Alonso-Gil / Marcos Davyt / Leonardo H F de Lima / Fanny Boissier / Sarada Das / Bojan Zagrovic / Joseph D Puglisi / Yaser Hashem / Zoya Ignatova /      要旨: The most abundant N-methyladenosine (mA) modification on mRNAs is installed non-stoichiometrically across transcripts, with 5' untranslated regions (5' UTRs) being the least conductive. 5' UTRs are ...The most abundant N-methyladenosine (mA) modification on mRNAs is installed non-stoichiometrically across transcripts, with 5' untranslated regions (5' UTRs) being the least conductive. 5' UTRs are essential for translation initiation, yet the molecular mechanisms orchestrated by mA remain poorly understood. Here, we combined structural, biochemical, and single-molecule approaches and show that at the most common position, a single mA does not affect translation yields, the kinetics of translation initiation complex assembly, or start codon recognition both under permissive growth and following exposure to oxidative stress. Cryoelectron microscopy (cryo-EM) structures of the late preinitiation complex reveal that mA purine ring established stacking interactions with an arginine side chain of the initiation factor eIF2α, although with only a marginal energy contribution, as estimated computationally. These findings provide molecular insights into mA interactions with the initiation complex and suggest that the subtle stabilization is unlikely to affect the translation dynamics under homeostatic conditions or stress. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_17330.map.gz emd_17330.map.gz | 170.6 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-17330-v30.xml emd-17330-v30.xml emd-17330.xml emd-17330.xml | 56.4 KB 56.4 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_17330_fsc.xml emd_17330_fsc.xml | 14.2 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_17330.png emd_17330.png | 32.1 KB | ||
| Filedesc metadata |  emd-17330.cif.gz emd-17330.cif.gz | 12.1 KB | ||
| その他 |  emd_17330_additional_1.map.gz emd_17330_additional_1.map.gz emd_17330_half_map_1.map.gz emd_17330_half_map_1.map.gz emd_17330_half_map_2.map.gz emd_17330_half_map_2.map.gz | 190.8 MB 168.3 MB 168.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17330 http://ftp.pdbj.org/pub/emdb/structures/EMD-17330 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17330 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17330 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_17330_validation.pdf.gz emd_17330_validation.pdf.gz | 1.1 MB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_17330_full_validation.pdf.gz emd_17330_full_validation.pdf.gz | 1.1 MB | 表示 | |
| XML形式データ |  emd_17330_validation.xml.gz emd_17330_validation.xml.gz | 21.8 KB | 表示 | |
| CIF形式データ |  emd_17330_validation.cif.gz emd_17330_validation.cif.gz | 28.4 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17330 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17330 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17330 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17330 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8p09MC  8p03C M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_17330.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_17330.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
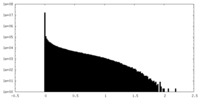
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: #1
| ファイル | emd_17330_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
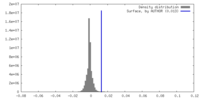
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_17330_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
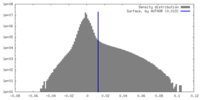
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_17330_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
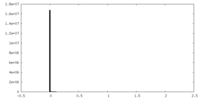
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : 48S late-stage initiation complex with non methylated mRNA
+超分子 #1: 48S late-stage initiation complex with non methylated mRNA
+分子 #1: initiator methionylated tRNA
+分子 #2: 18S ribosomal RNA
+分子 #3: mRNA
+分子 #4: Eukaryotic translation initiation factor 2 subunit 1
+分子 #5: 40S ribosomal protein SA
+分子 #6: ribosomal protein eS1
+分子 #7: 40S ribosomal protein S2
+分子 #8: Ribosomal protein S3
+分子 #9: 40S ribosomal protein S4
+分子 #10: Ribosomal protein S5
+分子 #11: 40S ribosomal protein S6
+分子 #12: ribosomal protein eS7
+分子 #13: 40S ribosomal protein S8
+分子 #14: 40S ribosomal protein S9
+分子 #15: 40S ribosomal protein eS10
+分子 #16: 40S ribosomal protein S11
+分子 #17: 40S ribosomal protein S12
+分子 #18: ribosomal protein uS15
+分子 #19: 40S ribosomal protein uS11
+分子 #20: 40S ribosomal protein uS19
+分子 #21: 40S ribosomal protein uS9
+分子 #22: 40S ribosomal protein eS17
+分子 #23: 40S ribosomal protein uS13
+分子 #24: 40S ribosomal protein eS19
+分子 #25: 40S ribosomal protein uS10
+分子 #26: 40S ribosomal protein S21
+分子 #27: Ribosomal protein S15a
+分子 #28: 40S ribosomal protein S23
+分子 #29: 40S ribosomal protein S24
+分子 #30: 40S ribosomal protein S26
+分子 #31: 40S ribosomal protein S27
+分子 #32: 40S ribosomal protein S28
+分子 #33: 40S ribosomal protein S29
+分子 #34: ribosomal protein eS31
+分子 #35: Ribosomal protein RACK1
+分子 #36: 40S ribosomal protein S30
+分子 #37: Eukaryotic translation initiation factor 4C
+分子 #38: ATP binding cassette subfamily E member 1
+分子 #39: 60S ribosomal protein L41
+分子 #40: 40S ribosomal protein S25
+分子 #41: MAGNESIUM ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 4 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TALOS ARCTICA |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 QUANTUM (4k x 4k) 平均電子線量: 45.0 e/Å2 |
| 電子線 | 加速電圧: 200 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: OTHER 最大 デフォーカス(公称値): 2.3000000000000003 µm 最小 デフォーカス(公称値): 0.3 µm |
| 実験機器 |  モデル: Talos Arctica / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)