+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Reconstruction of a 13 nm wide Abeta(1-40) amyloid fibril | |||||||||
 Map data Map data | This is a reconstruction of an Abeta(1-40) amyloid fibril | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Alzheimer's disease / amyloid / prion / protein folding | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 23.0 Å | |||||||||
 Authors Authors | Schmidt M / Sachse C / Richter W / Xu C / Fandrich M / Grigorieff N | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2009 Journal: Proc Natl Acad Sci U S A / Year: 2009Title: Comparison of Alzheimer Abeta(1-40) and Abeta(1-42) amyloid fibrils reveals similar protofilament structures. Authors: Matthias Schmidt / Carsten Sachse / Walter Richter / Chen Xu / Marcus Fändrich / Nikolaus Grigorieff /  Abstract: We performed mass-per-length (MPL) measurements and electron cryomicroscopy (cryo-EM) with 3D reconstruction on an Abeta(1-42) amyloid fibril morphology formed under physiological pH conditions. The ...We performed mass-per-length (MPL) measurements and electron cryomicroscopy (cryo-EM) with 3D reconstruction on an Abeta(1-42) amyloid fibril morphology formed under physiological pH conditions. The data show that the examined Abeta(1-42) fibril morphology has only one protofilament, although two protofilaments were observed with a previously studied Abeta(1-40) fibril. The latter fibril was resolved at 8 A resolution showing pairs of beta-sheets at the cores of the two protofilaments making up a fibril. Detailed comparison of the Abeta(1-42) and Abeta(1-40) fibril structures reveals that they share an axial twofold symmetry and a similar protofilament structure. Furthermore, the MPL data indicate that the protofilaments of the examined Abeta(1-40) and Abeta(1-42) fibrils have the same number of Abeta molecules per cross-beta repeat. Based on this data and the previously studied Abeta(1-40) fibril structure, we describe a model for the arrangement of peptides within the Abeta(1-42) fibril. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1650.map.gz emd_1650.map.gz | 58.5 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1650-v30.xml emd-1650-v30.xml emd-1650.xml emd-1650.xml | 9.8 KB 9.8 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-1650.tif EMD-1650.tif | 37.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1650 http://ftp.pdbj.org/pub/emdb/structures/EMD-1650 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1650 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1650 | HTTPS FTP |
-Validation report
| Summary document |  emd_1650_validation.pdf.gz emd_1650_validation.pdf.gz | 176 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1650_full_validation.pdf.gz emd_1650_full_validation.pdf.gz | 175.2 KB | Display | |
| Data in XML |  emd_1650_validation.xml.gz emd_1650_validation.xml.gz | 4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1650 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1650 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1650 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1650 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1650.map.gz / Format: CCP4 / Size: 62.5 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1650.map.gz / Format: CCP4 / Size: 62.5 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is a reconstruction of an Abeta(1-40) amyloid fibril | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.8 Å | ||||||||||||||||||||||||||||||||||||
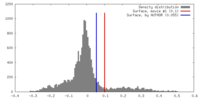
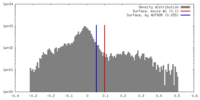
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Abeta(1-40) amyloid fibril
| Entire | Name: Abeta(1-40) amyloid fibril |
|---|---|
| Components |
|
-Supramolecule #1000: Abeta(1-40) amyloid fibril
| Supramolecule | Name: Abeta(1-40) amyloid fibril / type: sample / ID: 1000 / Oligomeric state: Cross-beta structure / Number unique components: 1 |
|---|
-Macromolecule #1: Abeta(1-40) peptide
| Macromolecule | Name: Abeta(1-40) peptide / type: protein_or_peptide / ID: 1 / Name.synonym: Alzheimer peptide / Oligomeric state: Cross-beta / Recombinant expression: Yes / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 4.33 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 8.7 / Details: 50 mM sodium borate |
| Grid | Details: 400 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 30 % / Chamber temperature: 90 K / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: Manual plunger (Brandeis) Method: One-sided blotting for 5 seconds before plunging |
| Details | Incubation for 4 days at 4 degC |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Temperature | Min: 90 K / Max: 90 K / Average: 90 K |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 4 / Average electron dose: 35 e/Å2 / Od range: 1.2 / Bits/pixel: 12 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 58090 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Fibrils were selected using BOXER |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Axial symmetry: C2 (2 fold cyclic) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 23.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider / Details: Final map was calculated from 4 fibrils |
| CTF correction | Details: Each particle |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)