+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Jumbo Phage phi-kp24 tail outer sheath | |||||||||
 Map data Map data | Jumbo phage phi-kp24 tail outer sheath | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Jumbo Phage / Klebsiella pneumoniae / tail / sheath / structural protein | |||||||||
| Function / homology | : / Bacteriophage phiKZ, gp29PR / Putative tail sheath protein / Putative virion structural protein Function and homology information Function and homology information | |||||||||
| Biological species |  Klebsiella phage vB_KpM_FBKp24 (virus) Klebsiella phage vB_KpM_FBKp24 (virus) | |||||||||
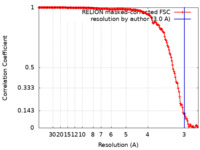
| Method | helical reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Ouyang R / Briegel A | |||||||||
| Funding support |  Netherlands, 1 items Netherlands, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: High-resolution reconstruction of a Jumbo-bacteriophage infecting capsulated bacteria using hyperbranched tail fibers. Authors: Ruochen Ouyang / Ana Rita Costa / C Keith Cassidy / Aleksandra Otwinowska / Vera C J Williams / Agnieszka Latka / Phill J Stansfeld / Zuzanna Drulis-Kawa / Yves Briers / Daniël M Pelt / ...Authors: Ruochen Ouyang / Ana Rita Costa / C Keith Cassidy / Aleksandra Otwinowska / Vera C J Williams / Agnieszka Latka / Phill J Stansfeld / Zuzanna Drulis-Kawa / Yves Briers / Daniël M Pelt / Stan J J Brouns / Ariane Briegel /      Abstract: The Klebsiella jumbo myophage ϕKp24 displays an unusually complex arrangement of tail fibers interacting with a host cell. In this study, we combine cryo-electron microscopy methods, protein ...The Klebsiella jumbo myophage ϕKp24 displays an unusually complex arrangement of tail fibers interacting with a host cell. In this study, we combine cryo-electron microscopy methods, protein structure prediction methods, molecular simulations, microbiological and machine learning approaches to explore the capsid, tail, and tail fibers of ϕKp24. We determine the structure of the capsid and tail at 4.1 Å and 3.0 Å resolution. We observe the tail fibers are branched and rearranged dramatically upon cell surface attachment. This complex configuration involves fourteen putative tail fibers with depolymerase activity that provide ϕKp24 with the ability to infect a broad panel of capsular polysaccharide (CPS) types of Klebsiella pneumoniae. Our study provides structural and functional insight into how ϕKp24 adapts to the variable surfaces of capsulated bacterial pathogens, which is useful for the development of phage therapy approaches against pan-drug resistant K. pneumoniae strains. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15669.map.gz emd_15669.map.gz | 35 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15669-v30.xml emd-15669-v30.xml emd-15669.xml emd-15669.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15669_fsc.xml emd_15669_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_15669.png emd_15669.png | 59.2 KB | ||
| Filedesc metadata |  emd-15669.cif.gz emd-15669.cif.gz | 5.9 KB | ||
| Others |  emd_15669_half_map_1.map.gz emd_15669_half_map_1.map.gz emd_15669_half_map_2.map.gz emd_15669_half_map_2.map.gz | 272.7 MB 272.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15669 http://ftp.pdbj.org/pub/emdb/structures/EMD-15669 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15669 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15669 | HTTPS FTP |
-Validation report
| Summary document |  emd_15669_validation.pdf.gz emd_15669_validation.pdf.gz | 825.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15669_full_validation.pdf.gz emd_15669_full_validation.pdf.gz | 824.7 KB | Display | |
| Data in XML |  emd_15669_validation.xml.gz emd_15669_validation.xml.gz | 23.3 KB | Display | |
| Data in CIF |  emd_15669_validation.cif.gz emd_15669_validation.cif.gz | 31.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15669 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15669 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15669 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15669 | HTTPS FTP |
-Related structure data
| Related structure data |  8au1MC  8bfkMC  8bflC  8bfpC  15776 C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15669.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15669.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Jumbo phage phi-kp24 tail outer sheath | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||
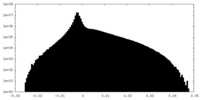
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: 2
| File | emd_15669_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
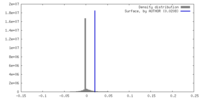
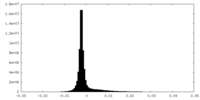
| Density Histograms |
-Half map: 1
| File | emd_15669_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Jumbo Phage phi-kp24 tail outer sheath
| Entire | Name: Jumbo Phage phi-kp24 tail outer sheath |
|---|---|
| Components |
|
-Supramolecule #1: Jumbo Phage phi-kp24 tail outer sheath
| Supramolecule | Name: Jumbo Phage phi-kp24 tail outer sheath / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: The outer sheath in extension of Klebsiella Phage phi-kp24 |
|---|---|
| Source (natural) | Organism:  Klebsiella phage vB_KpM_FBKp24 (virus) Klebsiella phage vB_KpM_FBKp24 (virus) |
-Macromolecule #1: Putative tail sheath protein
| Macromolecule | Name: Putative tail sheath protein / type: protein_or_peptide / ID: 1 / Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Klebsiella phage vB_KpM_FBKp24 (virus) Klebsiella phage vB_KpM_FBKp24 (virus) |
| Molecular weight | Theoretical: 76.4025 KDa |
| Sequence | String: MSEQITGSTP RIYYRGTKDS SVTRSTGSTT TLPLHRPLIM FFGQKGPTVP TWIDPVKFED IYGSETTNLS GVYCTHSTPF IKEAIAAGN QFMALRLEPS DIPDVATLGL SVDWVKTKID DYERNDDGTY KLDTNGDKIP LATQIDGIKF RFVLEKIETN E SGVSQYKK ...String: MSEQITGSTP RIYYRGTKDS SVTRSTGSTT TLPLHRPLIM FFGQKGPTVP TWIDPVKFED IYGSETTNLS GVYCTHSTPF IKEAIAAGN QFMALRLEPS DIPDVATLGL SVDWVKTKID DYERNDDGTY KLDTNGDKIP LATQIDGIKF RFVLEKIETN E SGVSQYKK RTAKAGTIGT EATPSTITPL ADFRCRFKSS LGANTALRIW APTINSAQAA DADLQARIKS FLYRFQILTR AD KASSPTI FETIYNEPSL SVGFGENLVD PQTEVVYDFV ERIDSRYNDE DPSTYLMSPL DTPYLYQANI DSVLTAIQEL EAP FDTVSA DEDDLYQINL FGAQTVEGVP YHAVQILGVL DGGVTLTETA TNYLQGGGDG TLGNDSFNAA AYAVLSNLSN NAAF NITNY ARYPFNAFWD SGFDLKTKQT IPQLIGLRAD TWIALSTQDI SSDFNSNEEE ESIALSLMSR VSAFPDSSDF GTPAF RGMI VGGAGYYTET TRKLPVPLTL DRFRAYCRYA GASDGVLKPE YAVDEGDARK VQVVKSINNL DKSWRVRRAQ WNNNLV YVE DYDTNSQFYP GQQSFYSEQG SVLKAAIVGL CVANLNRFAF EAWRDLTGTQ KLTDDQLIER SDDAVSTRGT GAFDDRL IF TPHSEITQAD KERGYSWSMR IDFGANAFRT VMDMSSVAYT REELANG UniProtKB: Putative tail sheath protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Support film - Material: CARBON / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: OTHER |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)