[English] 日本語
 Yorodumi
Yorodumi- EMDB-14539: CRYO-EM STRUCTURE OF SARS-COV-2 SPIKE : H11-H6 nanobody complex -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CRYO-EM STRUCTURE OF SARS-COV-2 SPIKE : H11-H6 nanobody complex | |||||||||
 Map data Map data | Sharpened map, output from postprocessing | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationvirion component / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion ...virion component / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / membrane fusion / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / symbiont-mediated suppression of host innate immune response / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |    Escherichia virus T4 Escherichia virus T4 | |||||||||
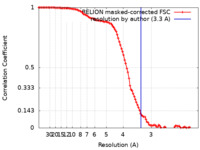
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Weckener M / Naismith JH | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Correlation between the binding affinity and the conformational entropy of nanobody SARS-CoV-2 spike protein complexes. Authors: Halina Mikolajek / Miriam Weckener / Z Faidon Brotzakis / Jiandong Huo / Evmorfia V Dalietou / Audrey Le Bas / Pietro Sormanni / Peter J Harrison / Philip N Ward / Steven Truong / Lucile ...Authors: Halina Mikolajek / Miriam Weckener / Z Faidon Brotzakis / Jiandong Huo / Evmorfia V Dalietou / Audrey Le Bas / Pietro Sormanni / Peter J Harrison / Philip N Ward / Steven Truong / Lucile Moynie / Daniel K Clare / Maud Dumoux / Joshua Dormon / Chelsea Norman / Naveed Hussain / Vinod Vogirala / Raymond J Owens / Michele Vendruscolo / James H Naismith /   Abstract: Camelid single-domain antibodies, also known as nanobodies, can be readily isolated from naïve libraries for specific targets but often bind too weakly to their targets to be immediately useful. ...Camelid single-domain antibodies, also known as nanobodies, can be readily isolated from naïve libraries for specific targets but often bind too weakly to their targets to be immediately useful. Laboratory-based genetic engineering methods to enhance their affinity, termed maturation, can deliver useful reagents for different areas of biology and potentially medicine. Using the receptor binding domain (RBD) of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike protein and a naïve library, we generated closely related nanobodies with micromolar to nanomolar binding affinities. By analyzing the structure-activity relationship using X-ray crystallography, cryoelectron microscopy, and biophysical methods, we observed that higher conformational entropy losses in the formation of the spike protein-nanobody complex are associated with tighter binding. To investigate this, we generated structural ensembles of the different complexes from electron microscopy maps and correlated the conformational fluctuations with binding affinity. This insight guided the engineering of a nanobody with improved affinity for the spike protein. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14539.map.gz emd_14539.map.gz | 12.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14539-v30.xml emd-14539-v30.xml emd-14539.xml emd-14539.xml | 29.4 KB 29.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14539_fsc.xml emd_14539_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_14539.png emd_14539.png | 129.8 KB | ||
| Masks |  emd_14539_msk_1.map emd_14539_msk_1.map | 103 MB |  Mask map Mask map | |
| Others |  emd_14539_additional_1.map.gz emd_14539_additional_1.map.gz emd_14539_additional_2.map.gz emd_14539_additional_2.map.gz emd_14539_half_map_1.map.gz emd_14539_half_map_1.map.gz emd_14539_half_map_2.map.gz emd_14539_half_map_2.map.gz | 6 MB 80.9 MB 80.9 MB 80.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14539 http://ftp.pdbj.org/pub/emdb/structures/EMD-14539 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14539 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14539 | HTTPS FTP |
-Validation report
| Summary document |  emd_14539_validation.pdf.gz emd_14539_validation.pdf.gz | 647.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14539_full_validation.pdf.gz emd_14539_full_validation.pdf.gz | 647.1 KB | Display | |
| Data in XML |  emd_14539_validation.xml.gz emd_14539_validation.xml.gz | 17.8 KB | Display | |
| Data in CIF |  emd_14539_validation.cif.gz emd_14539_validation.cif.gz | 23.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14539 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14539 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14539 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14539 | HTTPS FTP |
-Related structure data
| Related structure data |  7z7xMC  7z1aC  7z1bC  7z1cC  7z1dC  7z1eC  7z6vC  7z85C  7z86C  7z9qC  7z9rC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14539.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14539.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map, output from postprocessing | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14539_msk_1.map emd_14539_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
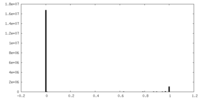
| Density Histograms |
-Additional map: Sharpened map, output from LocScale
| File | emd_14539_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map, output from LocScale | ||||||||||||
| Projections & Slices |
| ||||||||||||
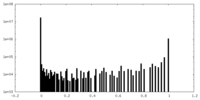
| Density Histograms |
-Additional map: Output from final 3D refinement
| File | emd_14539_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Output from final 3D refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
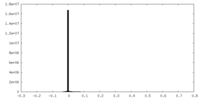
| Density Histograms |
-Half map: Half map from final 3D refinement
| File | emd_14539_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map from final 3D refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
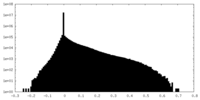
| Density Histograms |
-Half map: Half map from final 3D refinement
| File | emd_14539_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map from final 3D refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex between Spike and nanobody H11-H6
| Entire | Name: Complex between Spike and nanobody H11-H6 |
|---|---|
| Components |
|
-Supramolecule #1: Complex between Spike and nanobody H11-H6
| Supramolecule | Name: Complex between Spike and nanobody H11-H6 / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Molecular weight | Theoretical: 15 KDa |
-Supramolecule #2: Spike glycoprotein trimer with fibritin tag
| Supramolecule | Name: Spike glycoprotein trimer with fibritin tag / type: complex / Chimera: Yes / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK 293T Homo sapiens (human) / Recombinant strain: HEK 293T |
-Supramolecule #3: Nanobody H11-H6
| Supramolecule | Name: Nanobody H11-H6 / type: complex / Chimera: Yes / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Macromolecule #1: Spike glycoprotein,Fibritin
| Macromolecule | Name: Spike glycoprotein,Fibritin / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia virus T4 Escherichia virus T4 |
| Molecular weight | Theoretical: 139.877906 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF EYVSQPFLMD LEGKQGNFKN LREFVFKNID GYFKIYSKHT PINLVRDLPQ GFSALEPLVD LPIGINITRF QT LLALHRS YLTPGDSSSG WTAGAAAYYV GYLQPRTFLL KYNENGTITD AVDCALDPLS ETKCTLKSFT VEKGIYQTSN FRV QPTESI VRFPNITNLC PFGEVFNATR FASVYAWNRK RISNCVADYS VLYNSASFST FKCYGVSPTK LNDLCFTNVY ADSF VIRGD EVRQIAPGQT GKIADYNYKL PDDFTGCVIA WNSNNLDSKV GGNYNYLYRL FRKSNLKPFE RDISTEIYQA GSTPC NGVE GFNCYFPLQS YGFQPTNGVG YQPYRVVVLS FELLHAPATV CGPKKSTNLV KNKCVNFNFN GLTGTGVLTE SNKKFL PFQ QFGRDIADTT DAVRDPQTLE ILDITPCSFG GVSVITPGTN TSNQVAVLYQ DVNCTEVPVA IHADQLTPTW RVYSTGS NV FQTRAGCLIG AEHVNNSYEC DIPIGAGICA SYQTQTNSPG SASSVASQSI IAYTMSLGAE NSVAYSNNSI AIPTNFTI S VTTEILPVSM TKTSVDCTMY ICGDSTECSN LLLQYGSFCT QLNRALTGIA VEQDKNTQEV FAQVKQIYKT PPIKDFGGF NFSQILPDPS KPSKRSFIED LLFNKVTLAD AGFIKQYGDC LGDIAARDLI CAQKFNGLTV LPPLLTDEMI AQYTSALLAG TITSGWTFG AGAALQIPFA MQMAYRFNGI GVTQNVLYEN QKLIANQFNS AIGKIQDSLS STASALGKLQ DVVNQNAQAL N TLVKQLSS NFGAISSVLN DILSRLDPPE AEVQIDRLIT GRLQSLQTYV TQQLIRAAEI RASANLAATK MSECVLGQSK RV DFCGKGY HLMSFPQSAP HGVVFLHVTY VPAQEKNFTT APAICHDGKA HFPREGVFVS NGTHWFVTQR NFYEPQIITT DNT FVSGNC DVVIGIVNNT VYDPLQPELD SFKEELDKYF KNHTSPDVDL GDISGINASV VNIQKEIDRL NEVAKNLNES LIDL QELGK YEQGSGYIPE APRDGQAYVR KDGEWVLLST FLSLLNDIFE AQKIEWHEKH HHHHH |
-Macromolecule #2: Nanobody H11-H6
| Macromolecule | Name: Nanobody H11-H6 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.897658 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLVESGGG LMQAGGSLRL SCAVSGRTFS TAAMGWFRQA PGKEREFVAA IRWSGGSAYY ADSVKGRFTI SRDKAKNTVY LQMNSLKYE DTAVYYCAGS KITRSLLSDY ATWPYDYWGQ GTQVTVSSKH HHHHH |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 26 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: Homemade / Material: COPPER/RHODIUM / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE | |||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: SPOTITON Details: SPT Labtech prototype 300 mesh 1.2/2.0 nanowire grids with a highly reproduceable rectangular bar cross-section were used. The grids were glow-discharged on low for 120 s (Plasma Cleaner PDC- ...Details: SPT Labtech prototype 300 mesh 1.2/2.0 nanowire grids with a highly reproduceable rectangular bar cross-section were used. The grids were glow-discharged on low for 120 s (Plasma Cleaner PDC-002-CE, Harrick Plasma) or for 60-80 s, 12 mA (easiGlow, Pelco) to activate the nanowires. Approximately 6 nL of the complex were applied to the grids using a Chameleon EP system (SPT Labtech) at 79-81% relative humidity and ambient temperature.. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Details | EPU auto-function coma free correction |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 4 / Average exposure time: 3.0 sec. / Average electron dose: 50.0 e/Å2 / Details: The images were collected as 50 frame movies |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)