+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | 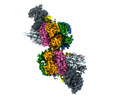 | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | S. cerevisiae CMGE dimer nucleating origin DNA melting | |||||||||||||||||||||||||||||||||
 マップデータ マップデータ | Consensus map of CMGE dimer model generated from EMD EMD-13978 | |||||||||||||||||||||||||||||||||
 試料 試料 |
| |||||||||||||||||||||||||||||||||
 キーワード キーワード | DNA replication / helicase / initiation / DNA origin / REPLICATION | |||||||||||||||||||||||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報DNA-templated DNA replication maintenance of fidelity / gene conversion / Unwinding of DNA / DNA replication initiation / epsilon DNA polymerase complex / MCM core complex / Assembly of the pre-replicative complex / Switching of origins to a post-replicative state / MCM complex binding / GINS complex ...DNA-templated DNA replication maintenance of fidelity / gene conversion / Unwinding of DNA / DNA replication initiation / epsilon DNA polymerase complex / MCM core complex / Assembly of the pre-replicative complex / Switching of origins to a post-replicative state / MCM complex binding / GINS complex / DNA strand elongation involved in mitotic DNA replication / nuclear DNA replication / mitotic DNA replication preinitiation complex assembly / premeiotic DNA replication / pre-replicative complex assembly involved in nuclear cell cycle DNA replication / mitotic DNA replication / SUMO binding / Activation of the pre-replicative complex / CMG complex / single-stranded 3'-5' DNA helicase activity / nuclear pre-replicative complex / single-stranded DNA 3'-5' DNA exonuclease activity / MCM complex / Activation of ATR in response to replication stress / DNA replication preinitiation complex / Termination of translesion DNA synthesis / replication fork protection complex / mitotic DNA replication checkpoint signaling / mitotic DNA replication initiation / double-strand break repair via break-induced replication / mitotic intra-S DNA damage checkpoint signaling / silent mating-type cassette heterochromatin formation / single-stranded DNA helicase activity / 加水分解酵素; エステル加水分解酵素; 5'-リン酸モノエステル産生エンドデオキシリボヌクレアーゼ / nucleotide-excision repair, DNA gap filling / mitotic sister chromatid cohesion / DNA replication proofreading / DNA strand elongation involved in DNA replication / leading strand elongation / DNA unwinding involved in DNA replication / nuclear replication fork / 3'-5' DNA helicase activity / DNA replication origin binding / Dual incision in TC-NER / subtelomeric heterochromatin formation / DNA replication initiation / error-prone translesion synthesis / DNA helicase activity / base-excision repair, gap-filling / helicase activity / replication fork / base-excision repair / DNA-templated DNA replication / double-strand break repair via nonhomologous end joining / double-strand break repair / mitotic cell cycle / single-stranded DNA binding / 4 iron, 4 sulfur cluster binding / double-stranded DNA binding / DNA helicase / cell cycle / chromosome, telomeric region / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / hydrolase activity / nucleotide binding / mRNA binding / chromatin binding / ATP hydrolysis activity / DNA binding / zinc ion binding / nucleoplasm / ATP binding / nucleus / metal ion binding / cytoplasm 類似検索 - 分子機能 | |||||||||||||||||||||||||||||||||
| 生物種 |  | |||||||||||||||||||||||||||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.4 Å | |||||||||||||||||||||||||||||||||
 データ登録者 データ登録者 | Lewis JS / Sousa JS | |||||||||||||||||||||||||||||||||
| 資金援助 | European Union,  フランス, フランス,  英国, 10件 英国, 10件
| |||||||||||||||||||||||||||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2022 ジャーナル: Nature / 年: 2022タイトル: Mechanism of replication origin melting nucleated by CMG helicase assembly. 著者: Jacob S Lewis / Marta H Gross / Joana Sousa / Sarah S Henrikus / Julia F Greiwe / Andrea Nans / John F X Diffley / Alessandro Costa /  要旨: The activation of eukaryotic origins of replication occurs in temporally separated steps to ensure that chromosomes are copied only once per cell cycle. First, the MCM helicase is loaded onto duplex ...The activation of eukaryotic origins of replication occurs in temporally separated steps to ensure that chromosomes are copied only once per cell cycle. First, the MCM helicase is loaded onto duplex DNA as an inactive double hexamer. Activation occurs after the recruitment of a set of firing factors that assemble two Cdc45-MCM-GINS (CMG) holo-helicases. CMG formation leads to the underwinding of DNA on the path to the establishment of the replication fork, but whether DNA becomes melted at this stage is unknown. Here we use cryo-electron microscopy to image ATP-dependent CMG assembly on a chromatinized origin, reconstituted in vitro with purified yeast proteins. We find that CMG formation disrupts the double hexamer interface and thereby exposes duplex DNA in between the two CMGs. The two helicases remain tethered, which gives rise to a splayed dimer, with implications for origin activation and replisome integrity. Inside each MCM ring, the double helix becomes untwisted and base pairing is broken. This comes as the result of ATP-triggered conformational changes in MCM that involve DNA stretching and protein-mediated stabilization of three orphan bases. Mcm2 pore-loop residues that engage DNA in our structure are dispensable for double hexamer loading and CMG formation, but are essential to untwist the DNA and promote replication. Our results explain how ATP binding nucleates origin DNA melting by the CMG and maintains replisome stability at initiation. | |||||||||||||||||||||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_14439.map.gz emd_14439.map.gz | 43.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-14439-v30.xml emd-14439-v30.xml emd-14439.xml emd-14439.xml | 40.2 KB 40.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_14439.png emd_14439.png | 83.3 KB | ||
| Filedesc metadata |  emd-14439.cif.gz emd-14439.cif.gz | 13.7 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14439 http://ftp.pdbj.org/pub/emdb/structures/EMD-14439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14439 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_14439_validation.pdf.gz emd_14439_validation.pdf.gz | 484.8 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_14439_full_validation.pdf.gz emd_14439_full_validation.pdf.gz | 484.3 KB | 表示 | |
| XML形式データ |  emd_14439_validation.xml.gz emd_14439_validation.xml.gz | 4.4 KB | 表示 | |
| CIF形式データ |  emd_14439_validation.cif.gz emd_14439_validation.cif.gz | 4.9 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14439 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14439 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14439 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14439 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_14439.map.gz / 形式: CCP4 / 大きさ: 60.2 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_14439.map.gz / 形式: CCP4 / 大きさ: 60.2 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Consensus map of CMGE dimer model generated from EMD EMD-13978 | ||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : S. cerevisiae CMGE dimer nucleating origin DNA melting
+超分子 #1: S. cerevisiae CMGE dimer nucleating origin DNA melting
+分子 #1: DNA replication licensing factor MCM2
+分子 #2: DNA replication licensing factor MCM3
+分子 #3: DNA replication licensing factor MCM4
+分子 #4: DNA helicase
+分子 #5: DNA replication licensing factor MCM6
+分子 #6: DNA replication licensing factor MCM7
+分子 #9: DNA replication complex GINS protein PSF3
+分子 #10: DNA replication complex GINS protein SLD5
+分子 #11: Cell division control protein 45
+分子 #12: DNA polymerase epsilon subunit B
+分子 #13: DNA replication complex GINS protein PSF1
+分子 #14: DNA replication complex GINS protein PSF2
+分子 #15: DNA polymerase epsilon catalytic subunit A
+分子 #7: DNA (53-MER)
+分子 #8: DNA (53-MER)
+分子 #16: ADENOSINE-5'-TRIPHOSPHATE
+分子 #17: ZINC ION
+分子 #18: MAGNESIUM ION
+分子 #19: ADENOSINE-5'-DIPHOSPHATE
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE / 装置: FEI VITROBOT MARK IV |
| 詳細 | four microlitres of sample was applied on a grid and incubated for 2 min at room temperature before blotting with filter paper for 5.5 s and plunge-freezing in liquid ethane. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / 撮影したグリッド数: 2 / 実像数: 65286 / 平均露光時間: 10.0 sec. / 平均電子線量: 1.6 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 50.0 µm / 照射モード: OTHER / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 4.4 µm / 最小 デフォーカス(公称値): 2.0 µm / 倍率(公称値): 130000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: NONE |
|---|---|
| 最終 再構成 | 解像度のタイプ: BY AUTHOR / 解像度: 3.4 Å / 解像度の算出法: FSC 0.143 CUT-OFF / ソフトウェア - 名称: RELION (ver. 3.1) / 使用した粒子像数: 71348 |
| 初期 角度割当 | タイプ: MAXIMUM LIKELIHOOD / ソフトウェア - 名称: RELION (ver. 3.1) |
| 最終 角度割当 | タイプ: MAXIMUM LIKELIHOOD / ソフトウェア - 名称: RELION (ver. 3.1) |
-原子モデル構築 1
| 初期モデル | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| 詳細 | One additional base pair has been built to connect the DNA molecules from the two individual symmetry expanded monomers. |
| 精密化 | プロトコル: RIGID BODY FIT |
| 得られたモデル |  PDB-7z13: |
 ムービー
ムービー コントローラー
コントローラー