[English] 日本語
 Yorodumi
Yorodumi- EMDB-14225: Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-14225 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ACE2 (Non-Uniform Refinement) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Spike / SARS-CoV-2 / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of amino acid transport / angiotensin-converting enzyme 2 / viral translation / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / regulation of systemic arterial blood pressure by renin-angiotensin / tryptophan transport / regulation of cardiac conduction ...positive regulation of amino acid transport / angiotensin-converting enzyme 2 / viral translation / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / regulation of systemic arterial blood pressure by renin-angiotensin / tryptophan transport / regulation of cardiac conduction / maternal process involved in female pregnancy / peptidyl-dipeptidase activity / regulation of vasoconstriction / transporter activator activity / Metabolism of Angiotensinogen to Angiotensins / carboxypeptidase activity / angiotensin maturation / Attachment and Entry / viral life cycle / receptor-mediated endocytosis of virus by host cell / metallocarboxypeptidase activity / positive regulation of cardiac muscle contraction / regulation of cytokine production / blood vessel diameter maintenance / negative regulation of smooth muscle cell proliferation / brush border membrane / negative regulation of ERK1 and ERK2 cascade / positive regulation of reactive oxygen species metabolic process / metallopeptidase activity / endocytic vesicle membrane / regulation of cell population proliferation / virus receptor activity / regulation of inflammatory response / endopeptidase activity / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / Potential therapeutics for SARS / structural constituent of virion / membrane fusion / Attachment and Entry / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / apical plasma membrane / host cell surface receptor binding / cilium / symbiont-mediated suppression of host innate immune response / membrane raft / endocytosis involved in viral entry into host cell / endoplasmic reticulum lumen / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / cell surface / negative regulation of transcription by RNA polymerase II / extracellular space / extracellular exosome / extracellular region / zinc ion binding / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
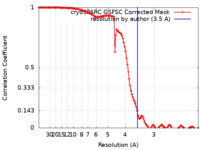
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Benton DJ / Wrobel AG | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Evolution of the SARS-CoV-2 spike protein in the human host. Authors: Antoni G Wrobel / Donald J Benton / Chloë Roustan / Annabel Borg / Saira Hussain / Stephen R Martin / Peter B Rosenthal / John J Skehel / Steven J Gamblin /  Abstract: Recently emerged variants of SARS-CoV-2 contain in their surface spike glycoproteins multiple substitutions associated with increased transmission and resistance to neutralising antibodies. We have ...Recently emerged variants of SARS-CoV-2 contain in their surface spike glycoproteins multiple substitutions associated with increased transmission and resistance to neutralising antibodies. We have examined the structure and receptor binding properties of spike proteins from the B.1.1.7 (Alpha) and B.1.351 (Beta) variants to better understand the evolution of the virus in humans. Spikes of both variants have the same mutation, N501Y, in the receptor-binding domains. This substitution confers tighter ACE2 binding, dependent on the common earlier substitution, D614G. Each variant spike has acquired other key changes in structure that likely impact virus pathogenesis. The spike from the Alpha variant is more stable against disruption upon binding ACE2 receptor than all other spikes studied. This feature is linked to the acquisition of a more basic substitution at the S1-S2 furin site (also observed for the variants of concern Delta, Kappa, and Omicron) which allows for near-complete cleavage. In the Beta variant spike, the presence of a new substitution, K417N (also observed in the Omicron variant), in combination with the D614G, stabilises a more open spike trimer, a conformation required for receptor binding. Our observations suggest ways these viruses have evolved to achieve greater transmissibility in humans. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14225.map.gz emd_14225.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14225-v30.xml emd-14225-v30.xml emd-14225.xml emd-14225.xml | 22 KB 22 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14225_fsc.xml emd_14225_fsc.xml | 13.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_14225.png emd_14225.png | 81.9 KB | ||
| Masks |  emd_14225_msk_1.map emd_14225_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-14225.cif.gz emd-14225.cif.gz | 7.6 KB | ||
| Others |  emd_14225_half_map_1.map.gz emd_14225_half_map_1.map.gz emd_14225_half_map_2.map.gz emd_14225_half_map_2.map.gz | 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14225 http://ftp.pdbj.org/pub/emdb/structures/EMD-14225 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14225 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14225 | HTTPS FTP |
-Related structure data
| Related structure data |  7r0zMC  7r10C  7r11C  7r12C  7r13C  7r14C  7r15C  7r16C  7r17C  7r18C  7r19C  7r1aC  7r1bC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14225.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14225.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.078 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_14225_msk_1.map emd_14225_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14225_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_14225_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ...
| Entire | Name: Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ACE2 (Non-Uniform Refinement) |
|---|---|
| Components |
|
-Supramolecule #1: Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ...
| Supramolecule | Name: Dissociated S1 domain of Alpha Variant SARS-CoV-2 Spike bound to ACE2 (Non-Uniform Refinement) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Molecular weight | Theoretical: 150 KDa |
-Supramolecule #2: Angiotensin-converting enzyme 2 (ACE2)
| Supramolecule | Name: Angiotensin-converting enzyme 2 (ACE2) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: SARS-CoV-2 Spike glycoprotein
| Supramolecule | Name: SARS-CoV-2 Spike glycoprotein / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Angiotensin-converting enzyme 2
| Macromolecule | Name: Angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: angiotensin-converting enzyme 2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 75.337422 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: METDTLLLWV LLLWVPGSTG STIEEQAKTF LDKFNHEAED LFYQSSLASW NYNTNITEEN VQNMNNAGDK WSAFLKEQST LAQMYPLQE IQNLTVKLQL QALQQNGSSV LSEDKSKRLN TILNTMSTIY STGKVCNPDN PQECLLLEPG LNEIMANSLD Y NERLWAWE ...String: METDTLLLWV LLLWVPGSTG STIEEQAKTF LDKFNHEAED LFYQSSLASW NYNTNITEEN VQNMNNAGDK WSAFLKEQST LAQMYPLQE IQNLTVKLQL QALQQNGSSV LSEDKSKRLN TILNTMSTIY STGKVCNPDN PQECLLLEPG LNEIMANSLD Y NERLWAWE SWRSEVGKQL RPLYEEYVVL KNEMARANHY EDYGDYWRGD YEVNGVDGYD YSRGQLIEDV EHTFEEIKPL YE HLHAYVR AKLMNAYPSY ISPIGCLPAH LLGDMWGRFW TNLYSLTVPF GQKPNIDVTD AMVDQAWDAQ RIFKEAEKFF VSV GLPNMT QGFWENSMLT DPGNVQKAVC HPTAWDLGKG DFRILMCTKV TMDDFLTAHH EMGHIQYDMA YAAQPFLLRN GANE GFHEA VGEIMSLSAA TPKHLKSIGL LSPDFQEDNE TEINFLLKQA LTIVGTLPFT YMLEKWRWMV FKGEIPKDQW MKKWW EMKR EIVGVVEPVP HDETYCDPAS LFHVSNDYSF IRYYTRTLYQ FQFQEALCQA AKHEGPLHKC DISNSTEAGQ KLFNML RLG KSEPWTLALE NVVGAKNMNV RPLLNYFEPL FTWLKDQNKN SFVGWSTDWS PYADDYKDDD DKWSHPQFEK GGGSGGG SG GSSAWSHPQF EK UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #2: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 141.981578 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GMFVFLVLLP LVSSQCVNLT TRTQLPPAYT NSFTRGVYYP DKVFRSSVLH STQDLFLPF FSNVTWFHAI SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV C EFQFCNDP ...String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GMFVFLVLLP LVSSQCVNLT TRTQLPPAYT NSFTRGVYYP DKVFRSSVLH STQDLFLPF FSNVTWFHAI SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV C EFQFCNDP FLGVYHKNNK SWMESEFRVY SSANNCTFEY VSQPFLMDLE GKQGNFKNLR EFVFKNIDGY FKIYSKHTPI NL VRDLPQG FSALEPLVDL PIGINITRFQ TLLALHRSYL TPGDSSSGWT AGAAAYYVGY LQPRTFLLKY NENGTITDAV DCA LDPLSE TKCTLKSFTV EKGIYQTSNF RVQPTESIVR FPNITNLCPF GEVFNATRFA SVYAWNRKRI SNCVADYSVL YNSA SFSTF KCYGVSPTKL NDLCFTNVYA DSFVIRGDEV RQIAPGQTGK IADYNYKLPD DFTGCVIAWN SNNLDSKVGG NYNYL YRLF RKSNLKPFER DISTEIYQAG STPCNGVEGF NCYFPLQSYG FQPTYGVGYQ PYRVVVLSFE LLHAPATVCG PKKSTN LVK NKCVNFNFNG LTGTGVLTES NKKFLPFQQF GRDIDDTTDA VRDPQTLEIL DITPCSFGGV SVITPGTNTS NQVAVLY QD VNCTEVPVAI HADQLTPTWR VYSTGSNVFQ TRAGCLIGAE HVNNSYECDI PIGAGICASY QTQTNSPSRA SSVASQSI I AYTMSLGAEN SVAYSNNSIA IPINFTISVT TEILPVSMTK TSVDCTMYIC GDSTECSNLL LQYGSFCTQL NRALTGIAV EQDKNTQEVF AQVKQIYKTP PIKDFGGFNF SQILPDPSKP SKRSFIEDLL FNKVTLADAG FIKQYGDCLG DIAARDLICA QKFNGLTVL PPLLTDEMIA QYTSALLAGT ITSGWTFGAG AALQIPFAMQ MAYRFNGIGV TQNVLYENQK LIANQFNSAI G KIQDSLSS TASALGKLQD VVNQNAQALN TLVKQLSSNF GAISSVLNDI LARLDPPEAE VQIDRLITGR LQSLQTYVTQ QL IRAAEIR ASANLAATKM SECVLGQSKR VDFCGKGYHL MSFPQSAPHG VVFLHVTYVP AQEKNFTTAP AICHDGKAHF PRE GVFVSN GTHWFVTQRN FYEPQIITTH NTFVSGNCDV VIGIVNNTVY DPLQPELDSF KEELDKYFKN HTSPDVDLGD ISGI NASVV NIQKEIDRLN EVAKNLNESL IDLQELGKYE QSGRENLYFQ GGGGSGYIPE APRDGQAYVR KDGEWVLLST FLGHH HHHH UniProtKB: Spike glycoprotein |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 8 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)