+ Open data
Open data
 Loading...
Loading...
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Large subunit of the Chlamydomonas reinhardtii mitoribosome | |||||||||||||||
 Map data Map data | Large subunit of the Chlamydomonas mitoribosome | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Mitochondria / mitoribosome / alga / RIBOSOME | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of mitochondrial mRNA stability / chloroplast rRNA processing / ribonucleoprotein granule / mitochondrial RNA processing / mitochondrial large ribosomal subunit / mitochondrial ribosome / mitochondrial translation / ribosome assembly / chloroplast / ribosomal large subunit assembly ...regulation of mitochondrial mRNA stability / chloroplast rRNA processing / ribonucleoprotein granule / mitochondrial RNA processing / mitochondrial large ribosomal subunit / mitochondrial ribosome / mitochondrial translation / ribosome assembly / chloroplast / ribosomal large subunit assembly / large ribosomal subunit / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / rRNA binding / negative regulation of translation / ribosome / structural constituent of ribosome / mitochondrial matrix / translation / mRNA binding / mitochondrion / RNA binding / cytoplasm Similarity search - Function Growth arrest and DNA damage-inducible proteins-interacting protein 1 domain superfamily / 39S ribosomal protein L46, mitochondrial / 39S ribosomal protein L43/54S ribosomal protein L51 / Ribosomal protein L47, mitochondrial / MRP-L47 superfamily, mitochondrial / Mitochondrial 39-S ribosomal protein L47 (MRP-L47) / Ribosomal protein L25, beta domain / Ribosomal protein L25, C-terminal / Ribosomal protein TL5, C-terminal domain / Ribosomal protein/NADH dehydrogenase domain ...Growth arrest and DNA damage-inducible proteins-interacting protein 1 domain superfamily / 39S ribosomal protein L46, mitochondrial / 39S ribosomal protein L43/54S ribosomal protein L51 / Ribosomal protein L47, mitochondrial / MRP-L47 superfamily, mitochondrial / Mitochondrial 39-S ribosomal protein L47 (MRP-L47) / Ribosomal protein L25, beta domain / Ribosomal protein L25, C-terminal / Ribosomal protein TL5, C-terminal domain / Ribosomal protein/NADH dehydrogenase domain / Mitochondrial ribosomal protein L51 / S25 / CI-B8 domain / Mitochondrial ribosomal protein L51 / S25 / CI-B8 domain / NUDIX hydrolase-like domain superfamily / Ribosomal protein L6, conserved site / Ribosomal protein L6 signature 1. / Ribosomal protein L9, C-terminal domain superfamily / Ribosomal protein L17 signature. / Ribosomal protein L28/L24 superfamily / Ribosomal protein L25/Gln-tRNA synthetase, anti-codon-binding domain superfamily / Ribosomal protein L9, N-terminal domain superfamily / Ribosomal protein L9 / Ribosomal protein L9, N-terminal / Ribosomal protein L9, N-terminal domain / Ribosomal protein L6, bacterial-type / Ribosomal protein L9/RNase H1, N-terminal / Ribosomal protein L20 signature. / Ribosomal protein L27, conserved site / Ribosomal protein L27 signature. / Ribosomal protein L22, bacterial/chloroplast-type / Ribosomal protein L2, bacterial/organellar-type / Ribosomal L28 family / Ribosomal protein L33 / Ribosomal protein L33 / Ribosomal protein L28/L24 / Ribosomal protein L33 superfamily / Ribosomal protein L30, bacterial-type / Ribosomal protein L16 / L28p-like / Ribosomal protein L20 / Ribosomal protein L20 / Ribosomal protein L20, C-terminal / Ribosomal protein L21 / Ribosomal protein L27 / Ribosomal L27 protein / Ribosomal protein L19 / Ribosomal protein L19 superfamily / Ribosomal protein L19 / Ribosomal proteins 50S L24/mitochondrial 39S L24 / Ribosomal protein L17 / Ribosomal protein L17 superfamily / Ribosomal protein L17 / Ribosomal protein L21-like / L21-like superfamily / Ribosomal prokaryotic L21 protein / Ribosomal L32p protein family / Ribosomal protein L32p / Ribosomal protein L24 / Ribosomal protein L34 / Ribosomal protein L34 / Ribosomal protein L13, bacterial-type / Ribosomal protein L3, bacterial/organelle-type / Ribosomal protein L15, bacterial-type / 50S ribosomal protein uL4 / Ribosomal protein L2 signature. / Ribosomal protein L2, conserved site / Ribosomal protein L5 domain superfamily / Ribosomal protein L10e/L16 / Ribosomal protein L10e/L16 superfamily / Ribosomal protein L16p/L10e / Ribosomal protein L6, alpha-beta domain / Ribosomal protein L6 / Ribosomal protein L6 / Ribosomal protein L6, alpha-beta domain superfamily / Ribosomal protein L2, domain 3 / Ribosomal protein L14P, conserved site / Ribosomal protein L14 signature. / Ribosomal Proteins L2, C-terminal domain / Ribosomal protein L2, C-terminal / Ribosomal Proteins L2, C-terminal domain / Ribosomal protein L24 signature. / Ribosomal Proteins L2, RNA binding domain / Ribosomal Proteins L2, RNA binding domain / Ribosomal protein L24/L26, conserved site / Ribosomal Proteins L2, RNA binding domain / Ribosomal protein L2 / KOW (Kyprides, Ouzounis, Woese) motif. / Ribosomal protein L13 / Ribosomal protein L13 / Ribosomal protein L13 superfamily / Ribosomal protein L30, ferredoxin-like fold domain / Ribosomal protein L30p/L7e / Ribosomal protein L23 / Ribosomal protein L15 / Ribosomal protein L25/L23 / Ribosomal protein L30, ferredoxin-like fold domain superfamily / Ribosomal proteins 50S-L15, 50S-L18e, 60S-L27A / Ribosomal protein L14p/L23e / Ribosomal protein L14P / Ribosomal protein L14 superfamily / Ribosomal protein L14p/L23e Similarity search - Domain/homology Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Ribosomal_L18e/L15P domain-containing protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / KOW domain-containing protein ...Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Ribosomal_L18e/L15P domain-containing protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / KOW domain-containing protein / Uncharacterized protein / Ribosomal_L2_C domain-containing protein / Ribosomal_L16 domain-containing protein / Uncharacterized protein / 50S ribosomal protein L9, chloroplastic / Uncharacterized protein / Ribosomal_L30 domain-containing protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Ribosomal_TL5_C domain-containing protein / Uncharacterized protein / Uncharacterized protein / 50S ribosomal protein L20 / Predicted protein / Predicted protein / Predicted protein / Mitochondrial ribosomal protein L19 / Uncharacterized protein / Mitochondrial ribosomal protein L17 / Predicted protein / Predicted protein / Predicted protein / Plastid ribosomal protein L6 / Predicted protein / Mitochondrial ribosomal protein L23 / Mitochondrial ribosomal protein L13 / Mitochondrial ribosomal protein L33 / Predicted protein / Mitochondrial ribosomal protein L17 Similarity search - Component | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||||||||
 Authors Authors | Waltz F / Soufari H / Hashem Y | |||||||||||||||
| Funding support | European Union,  France, 4 items France, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: How to build a ribosome from RNA fragments in Chlamydomonas mitochondria. Authors: Florent Waltz / Thalia Salinas-Giegé / Robert Englmeier / Herrade Meichel / Heddy Soufari / Lauriane Kuhn / Stefan Pfeffer / Friedrich Förster / Benjamin D Engel / Philippe Giegé / ...Authors: Florent Waltz / Thalia Salinas-Giegé / Robert Englmeier / Herrade Meichel / Heddy Soufari / Lauriane Kuhn / Stefan Pfeffer / Friedrich Förster / Benjamin D Engel / Philippe Giegé / Laurence Drouard / Yaser Hashem /    Abstract: Mitochondria are the powerhouse of eukaryotic cells. They possess their own gene expression machineries where highly divergent and specialized ribosomes, named hereafter mitoribosomes, translate the ...Mitochondria are the powerhouse of eukaryotic cells. They possess their own gene expression machineries where highly divergent and specialized ribosomes, named hereafter mitoribosomes, translate the few essential messenger RNAs still encoded by mitochondrial genomes. Here, we present a biochemical and structural characterization of the mitoribosome in the model green alga Chlamydomonas reinhardtii, as well as a functional study of some of its specific components. Single particle cryo-electron microscopy resolves how the Chlamydomonas mitoribosome is assembled from 13 rRNA fragments encoded by separate non-contiguous gene pieces. Additional proteins, mainly OPR, PPR and mTERF helical repeat proteins, are found in Chlamydomonas mitoribosome, revealing the structure of an OPR protein in complex with its RNA binding partner. Targeted amiRNA silencing indicates that these ribosomal proteins are required for mitoribosome integrity. Finally, we use cryo-electron tomography to show that Chlamydomonas mitoribosomes are attached to the inner mitochondrial membrane via two contact points mediated by Chlamydomonas-specific proteins. Our study expands our understanding of mitoribosome diversity and the various strategies these specialized molecular machines adopt for membrane tethering. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13480.map.gz emd_13480.map.gz | 394.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13480-v30.xml emd-13480-v30.xml emd-13480.xml emd-13480.xml | 80.8 KB 80.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13480.png emd_13480.png | 65.7 KB | ||
| Filedesc metadata |  emd-13480.cif.gz emd-13480.cif.gz | 16.3 KB | ||
| Others |  emd_13480_half_map_1.map.gz emd_13480_half_map_1.map.gz emd_13480_half_map_2.map.gz emd_13480_half_map_2.map.gz | 339.9 MB 340.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13480 http://ftp.pdbj.org/pub/emdb/structures/EMD-13480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13480 | HTTPS FTP |
-Related structure data
| Related structure data |  7pktMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13480.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13480.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Large subunit of the Chlamydomonas mitoribosome | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_13480_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
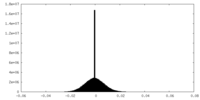
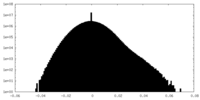
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_13480_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
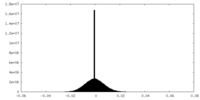
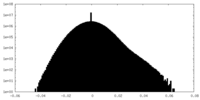
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Large subunit of the mitochondrial ribosome of Chlamydomonas rein...
| Entire | Name: Large subunit of the mitochondrial ribosome of Chlamydomonas reinhardtii |
|---|---|
| Components |
|
+Supramolecule #1: Large subunit of the mitochondrial ribosome of Chlamydomonas rein...
| Supramolecule | Name: Large subunit of the mitochondrial ribosome of Chlamydomonas reinhardtii type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#54 |
|---|---|
| Source (natural) | Organism:  |
+Macromolecule #1: Ribosomal_L2_C domain-containing protein
| Macromolecule | Name: Ribosomal_L2_C domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.087098 KDa |
| Sequence | String: MLRRVAQSCA TMLSKQGAGA MDMAQPSTSA PMFNLANEAL QAAGSGHGPA GTLLQFIRSK YTYKFKKAAP LPGLIYWKPS TPGTRHKIS IDYAGLGVYT GPPHPELSAH VAKTGGRNNT GRITTRHRGG GAPKVVRDVD FDRQRVDGLP GVVQRLEADP G RSGWLALV ...String: MLRRVAQSCA TMLSKQGAGA MDMAQPSTSA PMFNLANEAL QAAGSGHGPA GTLLQFIRSK YTYKFKKAAP LPGLIYWKPS TPGTRHKIS IDYAGLGVYT GPPHPELSAH VAKTGGRNNT GRITTRHRGG GAPKVVRDVD FDRQRVDGLP GVVQRLEADP G RSGWLALV KYSPEGQLPF YRYHLAPADV KAGDVLMSGH GAAIKPGNVL TLRDVPIGVP IHNIELVPGR GGQLARAAST SA VVTSKQD THAVVRLPSS ETRLIDLSCR ATVGQVSNHL QRSINHGKAG SRRNLGWRPS VRGIAMNPID HPHGGRTNGG RPS CTPWGV YTKGKRTRRR DASSNKFIIT RAGGQPIEKF VQAKKWRARA KLEAKQRAAA GGGGKRS UniProtKB: Ribosomal_L2_C domain-containing protein |
+Macromolecule #2: uL3m
| Macromolecule | Name: uL3m / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 44.936207 KDa |
| Sequence | String: MSATGSSCAL PTGIETSGGL PLQENSGDLA AASSAEASDA CRAASHGLAS TSLRCLATRE NHSFPDGWGT LCSQGLAASG LQLQQRRGY GYQPPKPRFP LPDSIASIVE EKTREHRRSA PAPPPAPKLE PRPMTPTSLR TGCIATKAGM TQEWDEHGVR V PLTVLWVD ...String: MSATGSSCAL PTGIETSGGL PLQENSGDLA AASSAEASDA CRAASHGLAS TSLRCLATRE NHSFPDGWGT LCSQGLAASG LQLQQRRGY GYQPPKPRFP LPDSIASIVE EKTREHRRSA PAPPPAPKLE PRPMTPTSLR TGCIATKAGM TQEWDEHGVR V PLTVLWVD ECQVVGLKTR PVHGYNALML GSGYKRQKCM SPSEAGFFLK AGVPFKKLVA EWQVSEDALV PVGTAIGAAH YV AGQRVDV TGWTKWKGFQ GVMRRWGFKG LPASHGVSLS HRAPGSIGNR QDPGKIWKGK KLPGCMGDER RTVHNCLVYK VDA ARNLVY LRGQVPGPVG RSVFLRDSRL ASPALRASWG LPFPTHVPSA EELKAAPAPG DVSVVPAPDG PGVTAWRNPT DPYL MYREE TDYMPVKWKK GE UniProtKB: Uncharacterized protein |
+Macromolecule #3: uL4m
| Macromolecule | Name: uL4m / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 45.910914 KDa |
| Sequence | String: MLAGALRGCA SESAFAWRQV VSAAAAAGSC SGSAGRLMVS SPCGVAQTSS PLQRWLFQGL RSSSTGAASI SGGLSEAGSL PPLVLRRVD DEALSPHPPT TDVGPLTVRY PFPIEYYKDR EAVIYSLDER PLGLAPLPGA AFNVPVRIDI LHRVVRYWRA K WQQGTHKA ...String: MLAGALRGCA SESAFAWRQV VSAAAAAGSC SGSAGRLMVS SPCGVAQTSS PLQRWLFQGL RSSSTGAASI SGGLSEAGSL PPLVLRRVD DEALSPHPPT TDVGPLTVRY PFPIEYYKDR EAVIYSLDER PLGLAPLPGA AFNVPVRIDI LHRVVRYWRA K WQQGTHKA KSRAEVSGGG KKPWNQKKTG RARQGSIRSP LWKGGGVSHA PRPRSHAHAL PRSTRLLGMR CALSAKINEG RF FVVDDLI NLRAAPLQ(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)SNKNP ARWSRHGLSP ADRPIREYGE LKRRLGALTE GSFGSSWLLV DSGEA GRDG GLRLRKLLKC SVVMEVVSPE ELTVYHVLKY HRLVVTRDAL QRISEALTRP HRVTKPVKHA WWARRRQAID AAVQEL TQA EAQA UniProtKB: Uncharacterized protein, Uncharacterized protein |
+Macromolecule #4: uL5m
| Macromolecule | Name: uL5m / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.054836 KDa |
| Sequence | String: MLKLQPRSWD ALPRLTAIEV SIPAIETQLE RDVVDKSELL LYALALEVLA GKPAGFTAPA NKALGTRATG VAVRLDAVTE PEAAHLFME KLVHVLLPNQ VGFEGVPPPM LVPPPRRSKA AEAAQARKAA LDHRKAPAKA HFTEIKVGNL LTYPDFEQNF S LFEPLRGM ...String: MLKLQPRSWD ALPRLTAIEV SIPAIETQLE RDVVDKSELL LYALALEVLA GKPAGFTAPA NKALGTRATG VAVRLDAVTE PEAAHLFME KLVHVLLPNQ VGFEGVPPPM LVPPPRRSKA AEAAQARKAA LDHRKAPAKA HFTEIKVGNL LTYPDFEQNF S LFEPLRGM RVRLVMEGAS AADCAALLGG MSLPVLSGAA AEAALAEITA EVARRARG UniProtKB: Uncharacterized protein |
+Macromolecule #5: 50S ribosomal protein L9, chloroplastic
| Macromolecule | Name: 50S ribosomal protein L9, chloroplastic / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.622402 KDa |
| Sequence | String: MQTAVAIGAR AGRGALLRSL LPCISGTIES ATSSQEAVER SGGLASTSST PAHGPSCACA RCFTSLAFST TRHASTAAAA KAKGKSSKS SAGRQAGTEA ADLDPGLIRV VLRKDVPRLG RRGEVVSVRR GLMRHSLYPA GEAAYATPDN IAKYGLADVE R GMDAAAEQ ...String: MQTAVAIGAR AGRGALLRSL LPCISGTIES ATSSQEAVER SGGLASTSST PAHGPSCACA RCFTSLAFST TRHASTAAAA KAKGKSSKS SAGRQAGTEA ADLDPGLIRV VLRKDVPRLG RRGEVVSVRR GLMRHSLYPA GEAAYATPDN IAKYGLADVE R GMDAAAEQ QQGGLEKLIK ALDTKPVVIT RRASDSDPES FAPGSAAVGA RAVAAAVESQ LGIRLDAASH LFLGEPIRSY GS YKVPLNL RLASEPQRQV ELTVHVDRRG KPQQPGATAA AGGGSSGEGG AGAAAGAAGG AGAAASS UniProtKB: 50S ribosomal protein L9, chloroplastic |
+Macromolecule #6: Mitochondrial ribosomal protein L13
| Macromolecule | Name: Mitochondrial ribosomal protein L13 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 29.714832 KDa |
| Sequence | String: MSRVTEAIKH LKDLQHIKLE GLRFRIIDAK GQVVGRLAEQ ISVILQGKDK PTYNPTKEQG DVVIVVNASH VELTHDKWNT KLYRWHTGY PGGLRSRPAG DQWEKDPRML LRNAVSGMLP KNKSREARME KLKIFPEADH PFQGFPLVPY VPPPRRLQDR G LGWPLPQG ...String: MSRVTEAIKH LKDLQHIKLE GLRFRIIDAK GQVVGRLAEQ ISVILQGKDK PTYNPTKEQG DVVIVVNASH VELTHDKWNT KLYRWHTGY PGGLRSRPAG DQWEKDPRML LRNAVSGMLP KNKSREARME KLKIFPEADH PFQGFPLVPY VPPPRRLQDR G LGWPLPQG FEPANLERYA FRLRTSPALQ AAAAPTVPAS SALAAAPQPS APVTGPGPAA SAPAAAAAAA GTAGASGSGS GR QQRQRPT VPIDDLLTEE ERAALAAVEG GKGKSGASSS UniProtKB: Mitochondrial ribosomal protein L13 |
+Macromolecule #7: uL14m
| Macromolecule | Name: uL14m / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.356652 KDa |
| Sequence | String: MIPKYSFLKI VDNSGAKLAQ VIGFYGHQGG YASVGDVVKV VVKEARGEKT TAGVMRKAVV VEQKYPFHRQ NGSHVQYLRN AAVLLSEKG QPIGNKVRSM LGVEFNKPRW AKLKSLSKRL F UniProtKB: Predicted protein |
+Macromolecule #8: Ribosomal_L18e/L15P domain-containing protein
| Macromolecule | Name: Ribosomal_L18e/L15P domain-containing protein / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 35.532332 KDa |
| Sequence | String: MSRSAAAGLL ARLGITCQRS CSVTFQIAST IAATASSAPS CSYTQHSPGA FASAGAPRGL ASLAFWTRRE AAASSSISDT LSSGAQPQQ QHLAASREAQ QARGFAAQAA GPDHLVGLGN LRDNPGATRQ RIRVGRGDGS RRGNYGGRGM KGQKSRGRGL H MLYDGGQL ...String: MSRSAAAGLL ARLGITCQRS CSVTFQIAST IAATASSAPS CSYTQHSPGA FASAGAPRGL ASLAFWTRRE AAASSSISDT LSSGAQPQQ QHLAASREAQ QARGFAAQAA GPDHLVGLGN LRDNPGATRQ RIRVGRGDGS RRGNYGGRGM KGQKSRGRGL H MLYDGGQL GLLKFPVTRQ RPSYEVLYYH LGLSRVAEFV QLGLLDAGRT ITMKDLYDSG CVTSNIKYGV LLYGKARLAV PL DLQVTAC DADTRAAVES AGGRVTRVYY TAEGLGAVLH PEKFTSRRLP LPLPAPRWHP RYDKKFDAIG QIPPATRPVA AAA LPAAAA HASGRVALAS A UniProtKB: Ribosomal_L18e/L15P domain-containing protein |
+Macromolecule #9: Ribosomal_L16 domain-containing protein
| Macromolecule | Name: Ribosomal_L16 domain-containing protein / type: protein_or_peptide / ID: 9 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.922502 KDa |
| Sequence | String: MVAEGSSAAT TSCSGRGLVT WTRQGPGASS ALGASLPGSS VPTFARLCPN LSVTAAVQLP GGVLLQRAGL HSLVASRAGS AAGAASAAR SFASPLGPLL APTAGASAAA AGASPGSSSG LLLGGWSACA GPVRWCQAGQ LSKQRVFKRE GINDVIMPNT T QLRYGTYG ...String: MVAEGSSAAT TSCSGRGLVT WTRQGPGASS ALGASLPGSS VPTFARLCPN LSVTAAVQLP GGVLLQRAGL HSLVASRAGS AAGAASAAR SFASPLGPLL APTAGASAAA AGASPGSSSG LLLGGWSACA GPVRWCQAGQ LSKQRVFKRE GINDVIMPNT T QLRYGTYG IRAMASKRVG AATIEAVRRT LRRKIKKTAR LWIRLSAVVP VTRKPLGIRM GKGKGVIDFY ATPVRPGQII FE MDRVPRK VALQALTAAQ HKFPVKLGFV EWS UniProtKB: Ribosomal_L16 domain-containing protein |
+Macromolecule #10: Mitochondrial ribosomal protein L17,bL17m
| Macromolecule | Name: Mitochondrial ribosomal protein L17,bL17m / type: protein_or_peptide / ID: 10 Details: The protein is a mixture of A0A2K3DXS2 and A8JH49,The protein is a mixture of A0A2K3DXS2 and A8JH49 Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.380727 KDa |
| Sequence | String: ARVRVRLGDW NRLNRKWQHR VSMMKTMVTQ LLEKERIRTT LAKAKTLQRY ADKVITLGKQ GTREAWSEAR GIVRTDRELH KLFTQVALR YRERPGGFTR LVRAGWRERD AAPVAFVELV DRPGELRPAN PALQRPNPLS VQAWIRQQLG QPAREPSPLL P PSVQQYLR EENRIQQGRA QPVTR UniProtKB: Mitochondrial ribosomal protein L17, Uncharacterized protein |
+Macromolecule #11: Mitochondrial ribosomal protein L19
| Macromolecule | Name: Mitochondrial ribosomal protein L19 / type: protein_or_peptide / ID: 11 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 33.412934 KDa |
| Sequence | String: MLGALLASLG ASGPRALSHY SASTSSLALQ SSIESIKETG AAYKALGTGW RFWSAGAGAS ARGDVNASSP AQAKSDESAA YASTAAPHE PAASPLNGDA QLSELLGDGG HPVASASGSS ARSGGGRSPP SGLPRLSLPQ RVLRDLRPPV QPYTLAPKLA Q RPTTTPAQ ...String: MLGALLASLG ASGPRALSHY SASTSSLALQ SSIESIKETG AAYKALGTGW RFWSAGAGAS ARGDVNASSP AQAKSDESAA YASTAAPHE PAASPLNGDA QLSELLGDGG HPVASASGSS ARSGGGRSPP SGLPRLSLPQ RVLRDLRPPV QPYTLAPKLA Q RPTTTPAQ YTPWTPTRAL QRRAHVSRRM GFLMHVLERE QMERVRAARP LPEFRAGDVL EVRMMVPEAE RKVVVYRGVC IA VYKKGIR SSFKIYNVFP DSGGVVQHIP LYMPDLVGVK VVGRIPASQE RLYYLLERET GTSDYTFQTN VSAAK UniProtKB: Mitochondrial ribosomal protein L19 |
+Macromolecule #12: 50S ribosomal protein L20
| Macromolecule | Name: 50S ribosomal protein L20 / type: protein_or_peptide / ID: 12 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.140527 KDa |
| Sequence | String: MPINRQKILQ LASSFRGRAR TCINIARLRV EKALQHAYVG RKLKKRHWRS VWIQRINAAS REHGMSYTGL ITGLKAENVA LNRKVLSEL AMSEPFSFKA LVEQVRHMRG LPPAGP UniProtKB: 50S ribosomal protein L20 |
+Macromolecule #13: bL21m
| Macromolecule | Name: bL21m / type: protein_or_peptide / ID: 13 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.170922 KDa |
| Sequence | String: MSLVRAALSQ LADSLSFGIL GVNRSCWGAS TSRATGLPSS ISGVRLLRTL AGTGDDQRAL PFNSRATVDE PFALPSTLAS STGCPTTPT PHAARLPALA ALHPPAGSLL PSLCQQLRGY KSNKRSLKPL NTAASWQAVQ PLLAEQRQRA AVEGVSPPVA D GARPPPQP ...String: MSLVRAALSQ LADSLSFGIL GVNRSCWGAS TSRATGLPSS ISGVRLLRTL AGTGDDQRAL PFNSRATVDE PFALPSTLAS STGCPTTPT PHAARLPALA ALHPPAGSLL PSLCQQLRGY KSNKRSLKPL NTAASWQAVQ PLLAEQRQRA AVEGVSPPVA D GARPPPQP GRRRWRPRLG PIVDTCLPPI PAVPLPTECR RVGVITGQYS LEPERTFAVV ELAGSQYKVT TDDVLFVNQL KG VAVNDVV ALERVLLLGS RAETVVGRPY VPGGVVLAAV EEHFRDGKVH VFKKRRRKRY RKYQAPRPHL TTLRILQVRG IDP APGDES ARLPELPVAL LDARYESVLR LTGRGEQQQQ QQQQQDEEQR QAAA UniProtKB: Uncharacterized protein |
+Macromolecule #14: uL22m
| Macromolecule | Name: uL22m / type: protein_or_peptide / ID: 14 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39.18007 KDa |
| Sequence | String: MSGAETHRLA LLASSRALIA RVQLGDAPAA LSQALSTLSS VIYASTSYGR SECSCSSSSG REVQPSATPA SPATAQTSRV APFLALQRP SAAFSRLGLS TWAPHASVSG WHGARQLHAG SAAQQAADVP GGSASGSGGK ESEGAAKPQE QTVTNPLAAA L ASAASGPV ...String: MSGAETHRLA LLASSRALIA RVQLGDAPAA LSQALSTLSS VIYASTSYGR SECSCSSSSG REVQPSATPA SPATAQTSRV APFLALQRP SAAFSRLGLS TWAPHASVSG WHGARQLHAG SAAQQAADVP GGSASGSGGK ESEGAAKPQE QTVTNPLAAA L ASAASGPV ASGSASAAGG LAQAAQAAAG RGRRQPRNWM WYDADDEREE RRKERQRLAW AMGPGGGEEV GAAHLMDVPQ SM KKMQRIV KLVRGLPYPD AVAQCSLVPH KAARYMLQAL EAAHADATEV KGLDAERLVV GTVFVTRGAY EPGISYHSKG RPG SKTFHR SHIRVVLNQA AERPAPFARI VAPLMSRRGL LGGSGGAGGA PRPRFAYRTE V UniProtKB: Uncharacterized protein |
+Macromolecule #15: Mitochondrial ribosomal protein L23
| Macromolecule | Name: Mitochondrial ribosomal protein L23 / type: protein_or_peptide / ID: 15 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.593826 KDa |
| Sequence | String: MRWQHSAAEQ TASTRAIKGS ADDTPSAAPE QETRLRFVSS EWRSGRPQPP ERRIPICFPN ITLQMMKLTP EQMDQVKETG WLREVAFKT TPDVTKLEIA AFLQSVYGMN VERVSTINYL GRRRLAISRN GKRLWWREDD WKKAYVIFRP PAGQEHLLTR G RRAGGDGD ...String: MRWQHSAAEQ TASTRAIKGS ADDTPSAAPE QETRLRFVSS EWRSGRPQPP ERRIPICFPN ITLQMMKLTP EQMDQVKETG WLREVAFKT TPDVTKLEIA AFLQSVYGMN VERVSTINYL GRRRLAISRN GKRLWWREDD WKKAYVIFRP PAGQEHLLTR G RRAGGDGD SGDEEQQRPV IEQIREAARA PRLPPRPYPW ERRPGGGRRG APGGEAEVPG GRSGDAAE UniProtKB: Mitochondrial ribosomal protein L23 |
+Macromolecule #16: KOW domain-containing protein,uL24m
| Macromolecule | Name: KOW domain-containing protein,uL24m / type: protein_or_peptide / ID: 16 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 31.1646 KDa |
| Sequence | String: MPQALYRSIQ PMFKTNRWRI CRDDLVYILS GPDKGKTGKV LEIIKDTRTP QVIVEGRNLR KKKVKTGTGP DDFFVVTMEA PLHYSQVQL VDPQTGKPMR VIFAYTADGT KVRSRKVPNP STADIIATPS EEDPDPRVGL TGPKDTSASL ARQATYTPSA G FPFRVRNL ...String: MPQALYRSIQ PMFKTNRWRI CRDDLVYILS GPDKGKTGKV LEIIKDTRTP QVIVEGRNLR KKKVKTGTGP DDFFVVTMEA PLHYSQVQL VDPQTGKPMR VIFAYTADGT KVRSRKVPNP STADIIATPS EEDPDPRVGL TGPKDTSASL ARQATYTPSA G FPFRVRNL ERIWAELQPD AD(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: KOW domain-containing protein |
+Macromolecule #17: Ribosomal_TL5_C domain-containing protein
| Macromolecule | Name: Ribosomal_TL5_C domain-containing protein / type: protein_or_peptide / ID: 17 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.730926 KDa |
| Sequence | String: MRCATGATAS LLPAARSWAV SMLTSGLDHG LGVSVLGGGP LPGLLARARH DSALAQRASE QVVPPEILRE LQQLEGSLND AARLDGRRR SSLRLSQSTL PLPLPTHTVE PGLPALDAHV RPRSATGGTA CKWLRRAGRV PGRVHSLPGA DARAGAAAGH S EDQGASSS ...String: MRCATGATAS LLPAARSWAV SMLTSGLDHG LGVSVLGGGP LPGLLARARH DSALAQRASE QVVPPEILRE LQQLEGSLND AARLDGRRR SSLRLSQSTL PLPLPTHTVE PGLPALDAHV RPRSATGGTA CKWLRRAGRV PGRVHSLPGA DARAGAAAGH S EDQGASSS SSNINNNNNK LLVHFSEGDV AKLIRTFGRN GCTARVLQLN LVAPDAAASD SGAEQAAQAD AGRSGEGTSG ST SGTQLGM LRVKPTRVFV NAVTQRVEGL DLIYCPPERQ VLVDVPVRLL NDDLAPGVKK GGWIALFKRT VRYKALGGAI PPY IEVNVR NMDLDQDMLV RDMPIPPGTK LYEKDYNAPV LRCTTDVGKD UniProtKB: Ribosomal_TL5_C domain-containing protein |
+Macromolecule #18: bL27m
| Macromolecule | Name: bL27m / type: protein_or_peptide / ID: 18 Details: N-ter is extend compared to A0A2K3E880,N-ter is extend compared to A0A2K3E880,N-ter is extend compared to A0A2K3E880,N-ter is extend compared to A0A2K3E880 Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.010873 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)MKLT AGELAFPGMI ILRQRGTKFH PGANVGIGRD HTIFATGVGK VRVDTQ PGP RGERRVVSVE PLPEALAAAS SPATPGAAAD ADSALRLRQV EAELVKRRAE VKRAMLQGRT PLEPALYFPL PRTADGQ LS WLERSVAKAP ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)MKLT AGELAFPGMI ILRQRGTKFH PGANVGIGRD HTIFATGVGK VRVDTQ PGP RGERRVVSVE PLPEALAAAS SPATPGAAAD ADSALRLRQV EAELVKRRAE VKRAMLQGRT PLEPALYFPL PRTADGQ LS WLERSVAKAP AVTPSAAAGK DAVPAKPKAA AKKAS UniProtKB: Uncharacterized protein |
+Macromolecule #19: bL28m
| Macromolecule | Name: bL28m / type: protein_or_peptide / ID: 19 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.000568 KDa |
| Sequence | String: MSSNVRQLIS RLIPPARKQF EHLQAHRRDV VWGDTQITLR VRQYPKSKDE RVSLVLPNWH RVRLWSEALG RKVELVMTKD TLRHIEDMG GLDAYLIKTP ESRLKSNPAS AVKWEVMSAL RRREAAAMMA RGGRGGAAGG DGSGAAGAAG KEST UniProtKB: Uncharacterized protein |
+Macromolecule #20: uL29m
| Macromolecule | Name: uL29m / type: protein_or_peptide / ID: 20 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.799233 KDa |
| Sequence | String: MRFDVENQGG LACVPKPCVS STGRGWKVEE LRLKSWQDLH KLWYVSLKER NLLLTEMGLK HIPKDAAHQV MRRIPVGAAS VEEDPHRLR YTEVQNTLRH IRQVLEERAA AELVPKLRRE MLEVINAR UniProtKB: Predicted protein |
+Macromolecule #21: Ribosomal_L30 domain-containing protein
| Macromolecule | Name: Ribosomal_L30 domain-containing protein / type: protein_or_peptide / ID: 21 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.367816 KDa |
| Sequence | String: MATEGSVQAV KSLFVTLRRG FAGTPWFHRR VLESLGLKYR HQCVEKPNNS SIRGMLSKVP HLVVIETDRM RYLRELKEHY QQLPREPWV LHHDPPLPAA TPSNPSSPLA STSSSDSDST ELLTPAPLPP SVPAQPRLTP FAASQLREHV EPVGLEGYRA N KQPPLPPR ...String: MATEGSVQAV KSLFVTLRRG FAGTPWFHRR VLESLGLKYR HQCVEKPNNS SIRGMLSKVP HLVVIETDRM RYLRELKEHY QQLPREPWV LHHDPPLPAA TPSNPSSPLA STSSSDSDST ELLTPAPLPP SVPAQPRLTP FAASQLREHV EPVGLEGYRA N KQPPLPPR KYFQEQSRRW RAKVLTERGV FSADACREHA EAIRPKSYRR RTGIRD UniProtKB: Ribosomal_L30 domain-containing protein |
+Macromolecule #22: bL32m
| Macromolecule | Name: bL32m / type: protein_or_peptide / ID: 22 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.370118 KDa |
| Sequence | String: MDSTSHPSTS YVVAPRLPAE EDDTFVPFSR PSPSPFGSGN SLLPQLCATP KKKLSLHRRG NRRVWYWLKR KTEVAKCSYC GALSGRNQM FSGAAGNSAC TGQCVEQAPT PPASSYPAEY KPPC UniProtKB: Predicted protein |
+Macromolecule #23: Mitochondrial ribosomal protein L33
| Macromolecule | Name: Mitochondrial ribosomal protein L33 / type: protein_or_peptide / ID: 23 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.864234 KDa |
| Sequence | String: MSNGGKKGVR LLVKLVSTAK TGFFYVTEKN PRNTPWKLKL MKFDPKVNKH VLFEEQKLK UniProtKB: Mitochondrial ribosomal protein L33 |
+Macromolecule #24: bL34m
| Macromolecule | Name: bL34m / type: protein_or_peptide / ID: 24 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 29.646521 KDa |
| Sequence | String: MASSSSVIAR AGQYGSLLLQ GPALSQATSN LGSLFSSTQA PPSADLSFSE RSSFALERLD TLTASSSGRQ HASGDAGLFV GFGGGAGGP LTAAHDSGLA PTICLWDSRL VSFDVSRLLA EAHRGRHQPQ TLSAQQPQLS HQELLGAHQH LPRLQMHLGG A DGVTPRLQ ...String: MASSSSVIAR AGQYGSLLLQ GPALSQATSN LGSLFSSTQA PPSADLSFSE RSSFALERLD TLTASSSGRQ HASGDAGLFV GFGGGAGGP LTAAHDSGLA PTICLWDSRL VSFDVSRLLA EAHRGRHQPQ TLSAQQPQLS HQELLGAHQH LPRLQMHLGG A DGVTPRLQ LHAPLTPPEL WAPHVLPGAP RPLQPHQPHQ PPAPAGLPAT PAPAPAPAPP APGGPAPAPL RCGNRGNTYQ PS RRKRVNR HGLEKRLSTP QGREVLLRRL RKGRWRITVD SFR UniProtKB: Uncharacterized protein |
+Macromolecule #25: bL35m
| Macromolecule | Name: bL35m / type: protein_or_peptide / ID: 25 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.493998 KDa |
| Sequence | String: MPWSLAHTPR GLFGAARPLV NPQIHAHDKY KKENPQPANS FLLGRFVTDR NGIIWHRQAN YRHARHAKSA SQLTRLKRWK PLAPAFAAK LRKLGFSERY WAAPDPQDVP GFHSPRGRVE RPRRSAVPDM DHTTGEPALR QSWQPPNRQR LR UniProtKB: Uncharacterized protein |
+Macromolecule #26: Plastid ribosomal protein L6
| Macromolecule | Name: Plastid ribosomal protein L6 / type: protein_or_peptide / ID: 26 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.275268 KDa |
| Sequence | String: MLTTRVKPFR AAAVSRSSRL CVVAKDSRIG RAPITVPKGV TVTLEGQLVR VKGPNGTLEQ TLSPLVKIEQ ADGKLKLFKL ADDRVAMSQ HGLNRSLVNN LVVGVSTGFE KRMEMVGTGY RAAVAGKDLT LNVGYSKPRV LAIPEGLKVV VEKNTTLVIS G ADKVKVGD ...String: MLTTRVKPFR AAAVSRSSRL CVVAKDSRIG RAPITVPKGV TVTLEGQLVR VKGPNGTLEQ TLSPLVKIEQ ADGKLKLFKL ADDRVAMSQ HGLNRSLVNN LVVGVSTGFE KRMEMVGTGY RAAVAGKDLT LNVGYSKPRV LAIPEGLKVV VEKNTTLVIS G ADKVKVGD FCATIRRQRP PEPYKGKGIR YAGEVIKLKE GKGAGGKKK UniProtKB: Plastid ribosomal protein L6 |
+Macromolecule #27: mL40
| Macromolecule | Name: mL40 / type: protein_or_peptide / ID: 27 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.749047 KDa |
| Sequence | String: MAGFTKATST KGAAMKRNKG AASGGVDEEN ARVKAMVAML QPPAPAEATP LTPAEQAAAQ QLQQQRVAEH QAWVKDMTTK FKLQQAALR ALPERLRVLA MQPDHTPYPL NRKFLFDSPP AAYRDTPAGK TASGASAAAE ASAGAGKGAE AASGGGKQAG G KQGGSRKQ QGGR UniProtKB: Uncharacterized protein |
+Macromolecule #28: mL41
| Macromolecule | Name: mL41 / type: protein_or_peptide / ID: 28 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.623615 KDa |
| Sequence | String: MTVRSIVVSL IRGANKASRQ HQGDIGREAV VDLIQQSAAK QSGIRKGWQV KAATWVKRVH VDRGDVKVGR LEGGEFQVLP HLRPRYFVP ADLDKFQLKP YVEVEKKVEA AKQ UniProtKB: Predicted protein |
+Macromolecule #29: mL43
| Macromolecule | Name: mL43 / type: protein_or_peptide / ID: 29 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.217593 KDa |
| Sequence | String: MSKYGVWMLE SLVIKYCDIG GSSRGMRLFL DEALPALRQQ NPQLGVQQVL QRFRHPKLVA VYRNGRTKPV CVKNLAPAEI MEHIGWLRN SHGRGQEYQV VRSRHLSRSP SIQGTWSVDT FASQLERVNE ARRAREAAQT QY UniProtKB: Predicted protein |
+Macromolecule #30: Mitochondrial ribosomal protein L17
| Macromolecule | Name: Mitochondrial ribosomal protein L17 / type: protein_or_peptide / ID: 30 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.320754 KDa |
| Sequence | String: MRSAGSLIAK SLFAVSTCGN RSCAGLLTRG YGAEAASATV NDGASTSGEG PIIAACVLQR LPLVRPRLTT GEQAQLALER QRALQTGEL KDYPQNMAVT GKDDAGSKQQ QQDLRAFDPV PATTAADSNG DVSTMRRRLA DSLYLVVRGG PAAGGGGGEW A FPQAVNTA ...String: MRSAGSLIAK SLFAVSTCGN RSCAGLLTRG YGAEAASATV NDGASTSGEG PIIAACVLQR LPLVRPRLTT GEQAQLALER QRALQTGEL KDYPQNMAVT GKDDAGSKQQ QQDLRAFDPV PATTAADSNG DVSTMRRRLA DSLYLVVRGG PAAGGGGGEW A FPQAVNTA EESISDTAQR ALYGAIGRSH PVYFIGNAPM GHVRALPAGT SFFMLAQVVD DPWDTQLLPG AAQELAWVTK QE LLGSYLS DARLRELVSK ML UniProtKB: Mitochondrial ribosomal protein L17 |
+Macromolecule #31: mL63/57/60
| Macromolecule | Name: mL63/57/60 / type: protein_or_peptide / ID: 31 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.645161 KDa |
| Sequence | String: MFFSRCVMVV FKTTGGRSWN PPSGLRPLSP AQRRNRTKNL ALTMKNMSIL KLAEANQPEV PVRLYKPLNF SRMQWMKKKL EETRAALGW DMEARALQEQ ARALRVGGGR QGAAGSLLPP AARAALQGSV GDK UniProtKB: Uncharacterized protein |
+Macromolecule #32: mL59/64
| Macromolecule | Name: mL59/64 / type: protein_or_peptide / ID: 32 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.372869 KDa |
| Sequence | String: MAHLLARLGA KALLPRRDGD KLLPPLIGAA EALRLREQFY AIGQQWPYEN IVPGKPPSPP GCKAYLERQK AKEAKRAARA KQVAEAMAN MPKTVADYKA SRRLDWSEVS ALDR(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) ...String: MAHLLARLGA KALLPRRDGD KLLPPLIGAA EALRLREQFY AIGQQWPYEN IVPGKPPSPP GCKAYLERQK AKEAKRAARA KQVAEAMAN MPKTVADYKA SRRLDWSEVS ALDR(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: Predicted protein |
+Macromolecule #33: mL80
| Macromolecule | Name: mL80 / type: protein_or_peptide / ID: 33 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.176438 KDa |
| Sequence | String: MHASVSGSLD TAPSSSSGTA LATASTSSAP LDLREQRHLY LDGTRTADPG EPRYTAPYWV PPSARAGIPN ILFSEPWPSH EEPQLRRQH AAMCLEALKR ADRPLTAEQV HEAVNSTAGY SASASAGDAG DSGAGADKPV LSTLAYTKKL LEHLRRTRFV Y GRKNPDSM ...String: MHASVSGSLD TAPSSSSGTA LATASTSSAP LDLREQRHLY LDGTRTADPG EPRYTAPYWV PPSARAGIPN ILFSEPWPSH EEPQLRRQH AAMCLEALKR ADRPLTAEQV HEAVNSTAGY SASASAGDAG DSGAGADKPV LSTLAYTKKL LEHLRRTRFV Y GRKNPDSM LSPGHPDHPR LYEALPFQAA RYGKPETLAA ADEAARAAAI AKAQKRLRNG KAPYPQHRRR ARFSIWQHEL AQ EALRELQ AK UniProtKB: Uncharacterized protein |
+Macromolecule #34: mL87
| Macromolecule | Name: mL87 / type: protein_or_peptide / ID: 34 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.535336 KDa |
| Sequence | String: MLALLAVRAR SPSLPSITLP ARLLSTQTSA SVSETYSNRP TSSAESTEAV SSSGQSASKW DWKWVLGKAS GRKPAITRPR RHQWHYCNP EYDPAAPLPE VLRSPFGPPG AERSHDWATY ARHLQLQPEN RRDLKRYRAR FVRFMQLREL DWREAFQRGV A EDSRVSNK ...String: MLALLAVRAR SPSLPSITLP ARLLSTQTSA SVSETYSNRP TSSAESTEAV SSSGQSASKW DWKWVLGKAS GRKPAITRPR RHQWHYCNP EYDPAAPLPE VLRSPFGPPG AERSHDWATY ARHLQLQPEN RRDLKRYRAR FVRFMQLREL DWREAFQRGV A EDSRVSNK VARAKAEAQR QDAWSDYKQA MWQRAQLADS HQSHGTGR UniProtKB: Predicted protein |
+Macromolecule #35: bL36m
| Macromolecule | Name: bL36m / type: protein_or_peptide / ID: 35 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 3.08179 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #36: mL113
| Macromolecule | Name: mL113 / type: protein_or_peptide / ID: 36 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 84.437922 KDa |
| Sequence | String: MRQLSLSLLA RLGRGSRGSL QPASSAINSG VDPGVLSGEE CSTSAPAGMP SWLRHSRRYA HQYNLIQPVD TNHINALLSS ATALEHAAV PVLRYSAWFD PEQVTRTMQR VPRMLQYQRR KGRRGAAYAS SPSSSADLAR SLLDALGSRL AALAPACSDQ Q LARALWAL ...String: MRQLSLSLLA RLGRGSRGSL QPASSAINSG VDPGVLSGEE CSTSAPAGMP SWLRHSRRYA HQYNLIQPVD TNHINALLSS ATALEHAAV PVLRYSAWFD PEQVTRTMQR VPRMLQYQRR KGRRGAAYAS SPSSSADLAR SLLDALGSRL AALAPACSDQ Q LARALWAL GAARHPHPQA LAAACEVLPQ RLKGASGAAA AAGAGSGAGG MAMTDLATAA WGLAAAASAG PQSVREPVRR AL QEVARHL VASRPADLSA TPALPQPSSP SSISSPSSGA VAPAADEVAA AASAAALAAD RPWLDPRSAV KLAWAFASCE VKD AAALDV VAEAAEARIA SQLQAHDPTT GPLTPRATYM YQTIRGWQAW PRPRPRVIRS AASAARGGRS RYLYDDRPRV VLRD FTAGS LAQLLAALAA AGHRHEGLMQ AAAAHLTASS GRSLRVDPHD LKRLAAAFAR LDLAAPAAAS GGAATAAALT ALLSA AQLS SLPAPLLARL AILAAESGVR RRSVYDRLVR QLMARAWVP(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) UniProtKB: Uncharacterized protein |
+Macromolecule #37: mL114
| Macromolecule | Name: mL114 / type: protein_or_peptide / ID: 37 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.757883 KDa |
| Sequence | String: MRRAVARCTA QIAAEACSVS SACSTSGRQV QEQLDNCLRW MTYARPRSRI DYYDPTRNVY KEFQRKWDRA NAAAAAAASQ AEAAAARPS HSAPPSAAPQ QQPYGGAGSY PFGSSLTTPC SSYGSGYGAG YGGTAAAGAD WPPGYEALLQ TYLEQELPLS A EEASTLVA ...String: MRRAVARCTA QIAAEACSVS SACSTSGRQV QEQLDNCLRW MTYARPRSRI DYYDPTRNVY KEFQRKWDRA NAAAAAAASQ AEAAAARPS HSAPPSAAPQ QQPYGGAGSY PFGSSLTTPC SSYGSGYGAG YGGTAAAGAD WPPGYEALLQ TYLEQELPLS A EEASTLVA AAKKGLLPGS RSLIRTRFMH IKDLEPRFPG FDARAAVLGE PRLLRHAADK VMRAMLVFQD HWPSHPVGPL MG RIGCPVI RDPAGVGHRL YALTRALKTD LHYELDPHRL TPESEGFLAS GVSPFELEAR VSALVTIFGR EGAGRLLDVS LDV LTYAPR DLDRAVLALR EVFSAAGDRG YGRHSLSTPE GAAAAAADRG YVTDLAVAWP GVLALPGRLG GADGVARLLA RVRR AGGAR YRGAVGRRAL LSEVLERPEL LQAAAEAAMR GEEEDEEEDV EGRAELEGKV UniProtKB: Uncharacterized protein |
+Macromolecule #38: mL115
| Macromolecule | Name: mL115 / type: protein_or_peptide / ID: 38 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 72.344484 KDa |
| Sequence | String: MSIALLCRRQ GRGLARALSL IESGASIERD VAITDENIRG SDAGACSTSR QCQPDTPIGG FRQWPRSLQL WRGFAVNNKP SLKYVPVPE SQHDDFTSEP TTYTQLLASI RAAKSLKRLE HLVDTYADRF DAVHVAAAVA RLPHLLKYRE ADLVDMSASA V VLPSGMTR ...String: MSIALLCRRQ GRGLARALSL IESGASIERD VAITDENIRG SDAGACSTSR QCQPDTPIGG FRQWPRSLQL WRGFAVNNKP SLKYVPVPE SQHDDFTSEP TTYTQLLASI RAAKSLKRLE HLVDTYADRF DAVHVAAAVA RLPHLLKYRE ADLVDMSASA V VLPSGMTR TRRKHGAQLR SGSAEVAARL AARVDAMLPQ HVAHFFPRQA ACSIWAFGEL RRHGVIERMD SLPQVLMSVT RG NLQPLRV HAAGVDFAQL LHGLAKLGHN DEPLLDALLP LVTERLGSMQ QRELQMTVWA LATLRRATPE LLDEVAQQLL STS TAFLLP SACASVFWSY AKTEGLARLS PGGEGGARVS RRRARVAAAR VKLFDSLAAT MMAQALLLAP QDVATTLWAC SVLG YHHSQ LPAVLGDAVL RALPNCSDAE VASVLESLAH LGYHHAPLMD AVAAGILAEP VVSTEPVNIA RVLYAYGVLA RRGPR DLQL VGTLAEALVR RLARVERLDT VALACRGLGA FAYDDQAVLA QVAARTEQLL QTSTTGLEQL QAVLRCLDAG GCSYFQ LAV AAARMLDDRL RAGACRSAPL AVEVLYYSAR QGV(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK) UniProtKB: Predicted protein |
+Macromolecule #39: mL116
| Macromolecule | Name: mL116 / type: protein_or_peptide / ID: 39 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.018789 KDa |
| Sequence | String: MRQQAGMLLG EAVASTSGRA SPALQLIIQR TLSLVASGIP SPRADVALQL STPHGGHINR MINTSESIVE LDSILYRFRK RLRPANIGA AAMRLEHLNR LERRTPYALR VQRVAAELQK YVATYTDRLA LTQAANVLRG LSAVRHRLPP ELVLRLAAGA V ADGGAALR ...String: MRQQAGMLLG EAVASTSGRA SPALQLIIQR TLSLVASGIP SPRADVALQL STPHGGHINR MINTSESIVE LDSILYRFRK RLRPANIGA AAMRLEHLNR LERRTPYALR VQRVAAELQK YVATYTDRLA LTQAANVLRG LSAVRHRLPP ELVLRLAAGA V ADGGAALR LAPDVDVRDL CFGLAGQGFN NTAFWARLCA AVLPRLRSFD PNTLPALVTA LQAAQQLPAP ASASASSGSA GS AVAAAAG GSTPQAAVAA EALRLLSRSE TLAALAPARL ADAASLLAGL GPALGVAVDA RLVEAVQTAT ARALPSLSPN QLP GLLLAV AALRRAAAPA EAAAAAAATA PQQQLPAALL ATALPHLSAG AVTMDLTAVM RAARLLAPHA AEPAAADTLV RLAR RTLLL LPAPGSSTGG PTGSASGSSS SNGEGLVTLS RVPRGGQAAG AVLAAAAPAG QLQGRTAGAV EGVARAFAAA APAVA PQPA LVGELAARLA AAGEAAAARG LLDEAQLASL GRSVEVLAAA GAAKSG UniProtKB: Uncharacterized protein |
+Macromolecule #40: mL117
| Macromolecule | Name: mL117 / type: protein_or_peptide / ID: 40 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 75.804086 KDa |
| Sequence | String: MLQALAGGAL GGLQTNGPAN LVGALGLLQR AAAAVVTGVP SSSSPVPPHA DRSLASLSAG AQSAAESACS HGGCGHDEAP CCSARSSSN SSSDAGAPRG LQQQLRSQQL QQHQHQQRRG IATSAGSALA YKFQSNVSPA SSRGSGRGSK VATRDNYQRW R ESGGDVRV ...String: MLQALAGGAL GGLQTNGPAN LVGALGLLQR AAAAVVTGVP SSSSPVPPHA DRSLASLSAG AQSAAESACS HGGCGHDEAP CCSARSSSN SSSDAGAPRG LQQQLRSQQL QQHQHQQRRG IATSAGSALA YKFQSNVSPA SSRGSGRGSK VATRDNYQRW R ESGGDVRV AQDILREAEG SGRGGGGGGG AGAARRGDSR LRRGADRPGG SGAGSGGGTG AGAGVVDVQD ELRAMVLGCR DL SELQVVV CECGADLNPF LVCAAAARLH KLKQATPPGA SPAALARRVG ESLMVLLQDR AAEAPLSQLA GAAHGLAEAG LAP GAALLE ALAARCEAAS PRGG(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) AAASELGAG MLRPLCDALT PRVPALSCAD VASLATGLAA ALGAAGGEAA AAAADGAPPL LSPSHFGSLP RLLSDLLLLR G PGQFGGRN FASVALALAL VTGGPAGAGG GGAAAAGSLP PAFWSKLAAV ALPEVPAMDA GSLSRLAGAF C(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: Uncharacterized protein, Uncharacterized protein |
+Macromolecule #41: mL118
| Macromolecule | Name: mL118 / type: protein_or_peptide / ID: 41 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 64.122902 KDa |
| Sequence | String: MAFLVAARRQ LKGGLSLAEV ALPVSVLGTC TRSNAVFTEA LAGCSGASLH CSAEQPLSRA TSYSDAGAGA AGVPACGSAS PRQPSPEPC SSSGSHATFN SLGRYRSQHV GISRGASALS ALAEASASPS PASNLPRGQH QHQQHHHQQL RTYHAWYYGS K LRNRAISQ ...String: MAFLVAARRQ LKGGLSLAEV ALPVSVLGTC TRSNAVFTEA LAGCSGASLH CSAEQPLSRA TSYSDAGAGA AGVPACGSAS PRQPSPEPC SSSGSHATFN SLGRYRSQHV GISRGASALS ALAEASASPS PASNLPRGQH QHQQHHHQQL RTYHAWYYGS K LRNRAISQ AESLEELGEM LVREGHRLDH VNLTALLAQL KRVARAAEEE AVAEATGGSS SSSSNSSSSG SSAVTAAAAA AA ARAVRVR VAELAAVAAR LVRRRAKWYD PRHAALAVAH TAALRHTDGR LLHDMTGRAL ARLDEAYSRD VLLLLRGLCA HQH MQQLAA ASSPPAVAAA VAPAVPAVAG AGKPYGGAPA VLLGGVKVFL TAKVPTGRMP PENLAGLLRH WRALAPPGRR LGPA VCGVV AADLQTRTAI YAPEPLAGVL ATLSAERHAL PPPLLDAAAE QFAAHALTHG SGAAAARFLA AVGAQLRLQQ QA (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: Uncharacterized protein |
+Macromolecule #42: mL119
| Macromolecule | Name: mL119 / type: protein_or_peptide / ID: 42 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.808164 KDa |
| Sequence | String: MATGAVAASA ADTASTSAPA PPSSSPFMPR WLRNLFPGGE HLPASPAMER QTSGASASSS GGSGPGSEAD DIKQLEEMRN MDMQGYVEY CKKMRGGAPP PRPRRPSVSP DHYDYRTMQD QRRIAFLRMQ QHEHIGSLVT KEESDLILAK REDVVKNRAL L QAIADRTG ...String: MATGAVAASA ADTASTSAPA PPSSSPFMPR WLRNLFPGGE HLPASPAMER QTSGASASSS GGSGPGSEAD DIKQLEEMRN MDMQGYVEY CKKMRGGAPP PRPRRPSVSP DHYDYRTMQD QRRIAFLRMQ QHEHIGSLVT KEESDLILAK REDVVKNRAL L QAIADRTG VYIDLEVKDC IEQFLETREN AGQMHRYATE FGMPLPKGSQ EQREMRRFMK RVEAEEKLAV ALEKRDLTSC SL RHKLSWA GPTALCDQTT LRYHECC(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)K REVVDKDRER QVHGMFKSRA EGILRKAAMD GVKPRLRDY UniProtKB: Uncharacterized protein, Uncharacterized protein |
+Macromolecule #43: Unk1
| Macromolecule | Name: Unk1 / type: protein_or_peptide / ID: 43 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 2.656265 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
+Macromolecule #44: PPR*
| Macromolecule | Name: PPR* / type: protein_or_peptide / ID: 44 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.656045 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
+Macromolecule #45: Unk2
| Macromolecule | Name: Unk2 / type: protein_or_peptide / ID: 45 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.188154 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) |
+Macromolecule #46: L1 rRNA
| Macromolecule | Name: L1 rRNA / type: rna / ID: 46 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 51.976762 KDa |
| Sequence | String: GGGAUUGCUG ACUGAUAGUG CAAACAGUAC UGUGAAGGAA CUAAUGAAAC UGGGCUAUAA GUGCGCAUUC CACCAAUACG UGAGCAUUC GCGCAUUACC AUUCUCCAGA GGAUUUAAAU GCACAAACCU AUAUGCCUUU UGGCAUAUGA UUACACUUCG U UA |
+Macromolecule #47: L2a rRNA
| Macromolecule | Name: L2a rRNA / type: rna / ID: 47 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.512189 KDa |
| Sequence | String: CAGCUAUAAU UUAGCGUACC UUUUGCAUUA UGGGUUAGCU ACUCAAUAGA CGUGAGUUAA GCAGAAAGCU UACUGAGAUU UUUUUA |
+Macromolecule #48: L3a rRNA (5S)
| Macromolecule | Name: L3a rRNA (5S) / type: rna / ID: 48 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.033252 KDa |
| Sequence | String: GGUGCUCGGU GAAACCGAGC AUCCAAUACU AAAGAAACUU UACUGGCUUA GUACUGGGAC CCCAUUUUUA GUAUA |
+Macromolecule #49: L3b rRNA
| Macromolecule | Name: L3b rRNA / type: rna / ID: 49 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 59.395223 KDa |
| Sequence | String: ACGUCUAUUG GACCCGAAAC CGAUCGAUCU AGCGACGUGC AGGUCUGACU UUAAUCAGGA CCGAACCCGU GUCUGUUGCA AGAGACUGG GAUGACACGU GGCUAGGGGU GAAAGGCCAA UCAAGUUCGG UAAUAGCUGG UUUUCUGCGA AAUCUAUUCA G GUAAAGAG UGUGUACUAA GACCAU |
+Macromolecule #50: L4 rRNA
| Macromolecule | Name: L4 rRNA / type: rna / ID: 50 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.253898 KDa |
| Sequence | String: AUACCGGUAG CGCAAGCGAA UGGUAAUGGA UCUAAUACAA CACAUUCAGA CAAUCGGCGG UAAGGUCAAU UGUCAAGAGG AGAACAGUC CAGAGGCGAG GCUAAGGUGC CUAACAAUU |
+Macromolecule #51: L5 rRNA
| Macromolecule | Name: L5 rRNA / type: rna / ID: 51 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.720117 KDa |
| Sequence | String: AAGGUAGAGU CUGCGUUGAA CACCAGAACA UAGGCUUGGA AGCAGCCACU GUGCUUGGAA AGCGUAAUAG CUCACUGGUC UAGCAGUCU CCGCUUAUAU AUGUCCGGCA CUAUUGCCGA AGCCCGCCUA UGACAAAUUU UUUUUUUUUU |
+Macromolecule #52: L6 rRNA
| Macromolecule | Name: L6 rRNA / type: rna / ID: 52 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.923266 KDa |
| Sequence | String: AGUCAUAGGU AGCAGAACGU UCUGGAACAA UUACGCUGAA AACAGUAACA GAAUUGAAGC UGCUAACAUA AGUAACACUA AAUGGAUAA CAUGGCCGUA AGCCCAAGGU UUUGCAUAAC AAACAGUGAA UACGGUU |
+Macromolecule #53: L7 rRNA
| Macromolecule | Name: L7 rRNA / type: rna / ID: 53 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 186.482641 KDa |
| Sequence | String: UACCGUACUG CAAACCGACU CAGGUGGGCU AGUAUAUAUA CUCAGGCCAU AGAAUCAACC AUGUUAAAGG AACUCGGCAA AAUGCACGC GUGACCUCGG AGGAAGCGUG GCUAAAAAGA CUCAGCAAAU AGCCGACAGC GACUGUUUAC CAAAAACACA G GAUAGUGC ...String: UACCGUACUG CAAACCGACU CAGGUGGGCU AGUAUAUAUA CUCAGGCCAU AGAAUCAACC AUGUUAAAGG AACUCGGCAA AAUGCACGC GUGACCUCGG AGGAAGCGUG GCUAAAAAGA CUCAGCAAAU AGCCGACAGC GACUGUUUAC CAAAAACACA G GAUAGUGC UAACAGACGU AAAUUAAGCU GCAAAAAGCG GUUUUAUUUU UGGAUGUAUA CUAUCCGACG CCAAACGGCG GC AGUAACU CUAACUGUCC UAAGGUAGCG AAAUUCCUUG ACUUGUAAUU GGCGUCCUGC AUGAAUGGCG GAACGACUGU CGG GCUGUC UCUAACAUGG AAUCUGUGAA UUUGAAAUCU UCGUGCAGAU GCGAAGGCGC UGGCAAGGGA CGAAGAGACC CUAU GCAGC UUCGCUGGGU GCUAAGAGUU ACUCUUAUCC UCAAGCUUUU CUGGGACGGA UGCUUCCUAA AUAGUAACGG GAGUG AUCA AUGGUACUGC UACUUAAAUA GAAAGCAGUG CUUGACUGUA AGAACCACAG UUCGAGCAGG GACGCAAGUC GGCAUU AAU GACCCAGGGU AACAU |
+Macromolecule #54: L8 rRNA
| Macromolecule | Name: L8 rRNA / type: rna / ID: 54 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 132.907203 KDa |
| Sequence | String: GUUACCCUGA UCAACGGACA AAAGCUACGC UAGGGAUAAC AGGCUUAUCC GCCUUUAGAG UUCCUAUCGA UGGGCGGGUU UGGCACCUC GAUGUCGGAU CAUCGCGUCC UGUGGCUGAA GGAGGUCGCA AGGGUUGGAU UGUUCGUCCA UUAAAGUGGU A CGUGAUCU ...String: GUUACCCUGA UCAACGGACA AAAGCUACGC UAGGGAUAAC AGGCUUAUCC GCCUUUAGAG UUCCUAUCGA UGGGCGGGUU UGGCACCUC GAUGUCGGAU CAUCGCGUCC UGUGGCUGAA GGAGGUCGCA AGGGUUGGAU UGUUCGUCCA UUAAAGUGGU A CGUGAUCU GAGUUCAGAC CGUCGUGAGA CAGUUCGGUU UCUAUCUUUG UCAGCGAAAG GCUUGUUUUG AAACGCCUAU GC ACAGUUA GCCGCUGGGG GCGAGUACGA AAGGACCAGA CCCCUAUAAA CCUCUGGUAC ACUUGUUGUC AUUUACACGG CAU AGCAAG AAAGCUAAGU UUUAAAAGGC UAGAUUGCCU ACAUUCAAUA ACACCGAAUG ACCCCGACUU ACAAAAAGUU UAGG ACUAA AUUUAUGA |
+Macromolecule #55: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 55 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
+Macromolecule #56: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 56 / Number of copies: 69 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
+Macromolecule #57: FE2/S2 (INORGANIC) CLUSTER
| Macromolecule | Name: FE2/S2 (INORGANIC) CLUSTER / type: ligand / ID: 57 / Number of copies: 1 / Formula: FES |
|---|---|
| Molecular weight | Theoretical: 175.82 Da |
| Chemical component information |  ChemComp-FES: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 7.6 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number grids imaged: 1 / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Number classes used: 2 / Resolution.type: BY AUTHOR / Resolution: 3.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 101291 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final 3D classification | Number classes: 6 / Software - Name: RELION |
+ About Yorodumi
About Yorodumi
-News
-Feb 9, 2022. New format data for meta-information of EMDB entries
New format data for meta-information of EMDB entries
- Version 3 of the EMDB header file is now the official format.
- The previous official version 1.9 will be removed from the archive.
Related info.: EMDB header
EMDB header
External links: wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
-Aug 12, 2020. Covid-19 info
Covid-19 info

URL: https://pdbjlc1.pdbj.org/emnavi/covid19.php
New page: Covid-19 featured information page in EM Navigator.
Related info.: Covid-19 info /
Covid-19 info /  Mar 5, 2020. Novel coronavirus structure data
Mar 5, 2020. Novel coronavirus structure data
+Mar 5, 2020. Novel coronavirus structure data
Novel coronavirus structure data
- International Committee on Taxonomy of Viruses (ICTV) defined the short name of the 2019 coronavirus as "SARS-CoV-2".
- In the structure databanks used in Yorodumi, some data are registered as the other names, "COVID-19 virus" and "2019-nCoV". Here are the details of the virus and the list of structure data.
Related info.: Yorodumi Speices /
Yorodumi Speices /  Aug 12, 2020. Covid-19 info
Aug 12, 2020. Covid-19 info
External links: COVID-19 featured content - PDBj /
COVID-19 featured content - PDBj /  Molecule of the Month (242):Coronavirus Proteases
Molecule of the Month (242):Coronavirus Proteases
+Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
EMDB accession codes are about to change! (news from PDBe EMDB page)
- The allocation of 4 digits for EMDB accession codes will soon come to an end. Whilst these codes will remain in use, new EMDB accession codes will include an additional digit and will expand incrementally as the available range of codes is exhausted. The current 4-digit format prefixed with “EMD-” (i.e. EMD-XXXX) will advance to a 5-digit format (i.e. EMD-XXXXX), and so on. It is currently estimated that the 4-digit codes will be depleted around Spring 2019, at which point the 5-digit format will come into force.
- The EM Navigator/Yorodumi systems omit the EMD- prefix.
Related info.: Q: What is EMD? /
Q: What is EMD? /  ID/Accession-code notation in Yorodumi/EM Navigator
ID/Accession-code notation in Yorodumi/EM Navigator
External links: EMDB Accession Codes are Changing Soon! /
EMDB Accession Codes are Changing Soon! /  Contact to PDBj
Contact to PDBj
+Jul 12, 2017. Major update of PDB
Major update of PDB
- wwPDB released updated PDB data conforming to the new PDBx/mmCIF dictionary.
- This is a major update changing the version number from 4 to 5, and with Remediation, in which all the entries are updated.
- In this update, many items about electron microscopy experimental information are reorganized (e.g. em_software).
- Now, EM Navigator and Yorodumi are based on the updated data.
External links: wwPDB Remediation /
wwPDB Remediation /  Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
-Yorodumi
Thousand views of thousand structures
- Yorodumi is a browser for structure data from EMDB, PDB, SASBDB, etc.
- This page is also the successor to EM Navigator detail page, and also detail information page/front-end page for Omokage search.
- The word "yorodu" (or yorozu) is an old Japanese word meaning "ten thousand". "mi" (miru) is to see.
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  Yorodumi Search /
Yorodumi Search /  Aug 31, 2016. New EM Navigator & Yorodumi /
Aug 31, 2016. New EM Navigator & Yorodumi /  Yorodumi Papers /
Yorodumi Papers /  Jmol/JSmol /
Jmol/JSmol /  Function and homology information /
Function and homology information /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X