[English] 日本語
 Yorodumi
Yorodumi- EMDB-1316: High-resolution electron microscopy of helical specimens: a fresh... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1316 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
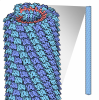
| Title | High-resolution electron microscopy of helical specimens: a fresh look at tobacco mosaic virus. | |||||||||
 Map data Map data | A 69 Angstrom section of the virus. The complete virus contains contains approximately 44 repeats of this 69 Angstrom section. The section contains 49 asymmetric units of the helical virus.Cube of 241 x 241 x 241 Angstrom | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | Tobacco mosaic virus-like, coat protein / Tobacco mosaic virus-like, coat protein superfamily / Virus coat protein (TMV like) / helical viral capsid / structural molecule activity /  Capsid protein Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |    Tobacco mosaic virus Tobacco mosaic virus | |||||||||
| Method | helical reconstruction /  cryo EM / Resolution: 4.4 Å cryo EM / Resolution: 4.4 Å | |||||||||
 Authors Authors | Sachse C / Chen JZ / Coureux PD / Stroupe ME / Fandrich M / Grigorieff N | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2007 Journal: J Mol Biol / Year: 2007Title: High-resolution electron microscopy of helical specimens: a fresh look at tobacco mosaic virus. Authors: Carsten Sachse / James Z Chen / Pierre-Damien Coureux / M Elizabeth Stroupe / Marcus Fändrich / Nikolaus Grigorieff /  Abstract: The treatment of helical objects as a string of single particles has become an established technique to resolve their three-dimensional (3D) structure using electron cryo-microscopy. It can be ...The treatment of helical objects as a string of single particles has become an established technique to resolve their three-dimensional (3D) structure using electron cryo-microscopy. It can be applied to a wide range of helical particles such as viruses, microtubules and helical filaments. We have made improvements to this approach using Tobacco Mosaic Virus (TMV) as a test specimen and obtained a map from 210,000 asymmetric units at a resolution better than 5 A. This was made possible by performing a full correction of the contrast transfer function of the microscope. Alignment of helical segments was helped by constraints derived from the helical symmetry of the virus. Furthermore, symmetrization was implemented by multiple inclusions of symmetry-related views in the 3D reconstruction. We used the density map to build an atomic model of TMV. The model was refined using a real-space refinement strategy that accommodates multiple conformers. The atomic model shows significant deviations from the deposited model for the helical form of TMV at the lower-radius region (residues 88 to 109). This region appears more ordered with well-defined secondary structure, compared with the earlier helical structure. The RNA phosphate backbone is sandwiched between two arginine side-chains, stabilizing the interaction between RNA and coat protein. A cluster of two or three carboxylates is buried in a hydrophobic environment isolating it from neighboring subunits. These carboxylates may represent the so-called Caspar carboxylates that form a metastable switch for viral disassembly. Overall, the observed differences suggest that the new model represents a different, more stable state of the virus, compared with the earlier published model. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1316.map.gz emd_1316.map.gz | 8.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1316-v30.xml emd-1316-v30.xml emd-1316.xml emd-1316.xml | 11.6 KB 11.6 KB | Display Display |  EMDB header EMDB header |
| Images |  1316.gif 1316.gif | 60 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1316 http://ftp.pdbj.org/pub/emdb/structures/EMD-1316 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1316 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1316 | HTTPS FTP |
-Related structure data
| Related structure data |  2om3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1316.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1316.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | A 69 Angstrom section of the virus. The complete virus contains contains approximately 44 repeats of this 69 Angstrom section. The section contains 49 asymmetric units of the helical virus.Cube of 241 x 241 x 241 Angstrom | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.163 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Tobacco Mosaic Virus
| Entire | Name:    Tobacco Mosaic Virus Tobacco Mosaic Virus |
|---|---|
| Components |
|
-Supramolecule #1000: Tobacco Mosaic Virus
| Supramolecule | Name: Tobacco Mosaic Virus / type: sample / ID: 1000 / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 37.5 MDa |
-Supramolecule #1: Tobacco mosaic virus
| Supramolecule | Name: Tobacco mosaic virus / type: virus / ID: 1 / Name.synonym: Virus / NCBI-ID: 12242 / Sci species name: Tobacco mosaic virus / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: Virus |
|---|---|
| Host (natural) | Organism:   Nicotiana tabacum (common tobacco) / synonym: PLANTAE(HIGHER PLANTS) Nicotiana tabacum (common tobacco) / synonym: PLANTAE(HIGHER PLANTS) |
| Molecular weight | Experimental: 37.5 MDa |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: Phosphate (5mM EDTA) |
| Grid | Details: Quantifoil grid |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: self-built model / Method: Blot for 2 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 60190 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 4.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 59000 Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 4.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder Specimen holder model: GATAN LIQUID NITROGEN |
| Temperature | Min: 93 K / Max: 93 K / Average: 93 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 100,000 times magnification |
| Date | Nov 22, 2005 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7.0 µm / Number real images: 6 / Average electron dose: 15 e/Å2 / Bits/pixel: 8 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: CTFFIND and CTFTILT phase-correction and amplitude-weighting of each paritcle after Grigorieff 1998 |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 4.4 Å / Resolution method: OTHER / Software - Name: Spider Details: Final map was calculated from the central 241x241x241 cube. |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - #0 - Chain ID: A / Chain - #1 - Chain ID: R |
|---|---|
| Software | Name:  XPLOR XPLOR |
| Details | Protocol: Torsion angle dynamics. 2TMV pdb code structure was adjusted to fit the density for residues 88-109 and refined |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 90 Target criteria: minimization of of least-square difference between observed and calculated densities |
| Output model |  PDB-2om3: |
 Movie
Movie Controller
Controller