[English] 日本語
 Yorodumi
Yorodumi- EMDB-12303: Mammalian ribosome nascent chain complex with SRP and SRP recepto... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12303 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mammalian ribosome nascent chain complex with SRP and SRP receptor in early state A | |||||||||
 Map data Map data | complex of RNC with SRPG226E and SRP receptor | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Signal recognition particle / protein targeting to the ER membrane / RIBOSOME | |||||||||
| Function / homology |  Function and homology information Function and homology informationPINK1-PRKN Mediated Mitophagy / signal recognition particle receptor complex / SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition / endoplasmic reticulum signal peptide binding / signal recognition particle, endoplasmic reticulum targeting / Translesion synthesis by REV1 / Recognition of DNA damage by PCNA-containing replication complex / Translesion Synthesis by POLH / Downregulation of ERBB4 signaling / Spry regulation of FGF signaling ...PINK1-PRKN Mediated Mitophagy / signal recognition particle receptor complex / SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition / endoplasmic reticulum signal peptide binding / signal recognition particle, endoplasmic reticulum targeting / Translesion synthesis by REV1 / Recognition of DNA damage by PCNA-containing replication complex / Translesion Synthesis by POLH / Downregulation of ERBB4 signaling / Spry regulation of FGF signaling / Downregulation of ERBB2:ERBB3 signaling / NOD1/2 Signaling Pathway / APC/C:Cdc20 mediated degradation of Cyclin B / SCF-beta-TrCP mediated degradation of Emi1 / APC-Cdc20 mediated degradation of Nek2A / EGFR downregulation / TCF dependent signaling in response to WNT / NRIF signals cell death from the nucleus / p75NTR recruits signalling complexes / NF-kB is activated and signals survival / Activated NOTCH1 Transmits Signal to the Nucleus / Downregulation of TGF-beta receptor signaling / TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) / Downregulation of SMAD2/3:SMAD4 transcriptional activity / SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription / Senescence-Associated Secretory Phenotype (SASP) / Regulation of innate immune responses to cytosolic DNA / activated TAK1 mediates p38 MAPK activation / JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 / Regulation of FZD by ubiquitination / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / Regulation of TNFR1 signaling / TNFR1-induced NF-kappa-B signaling pathway / Translesion synthesis by POLK / Translesion synthesis by POLI / Regulation of necroptotic cell death / MAP3K8 (TPL2)-dependent MAPK1/3 activation / HDR through Homologous Recombination (HRR) / Josephin domain DUBs / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / Processing of DNA double-strand break ends / Formation of Incision Complex in GG-NER / Gap-filling DNA repair synthesis and ligation in GG-NER / Dual Incision in GG-NER / Fanconi Anemia Pathway / Regulation of TP53 Activity through Phosphorylation / Regulation of TP53 Degradation / Regulation of TP53 Activity through Methylation / Negative regulation of MET activity / Cyclin D associated events in G1 / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / Downregulation of ERBB2 signaling / E3 ubiquitin ligases ubiquitinate target proteins / Regulation of PTEN localization / ER Quality Control Compartment (ERQC) / Regulation of expression of SLITs and ROBOs / Interferon alpha/beta signaling / Endosomal Sorting Complex Required For Transport (ESCRT) / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / IKK complex recruitment mediated by RIP1 / IRAK2 mediated activation of TAK1 complex / TRAF6-mediated induction of TAK1 complex within TLR4 complex / Alpha-protein kinase 1 signaling pathway / RAS processing / Pexophagy / Inactivation of CSF3 (G-CSF) signaling / Negative regulation of FLT3 / Regulation of BACH1 activity / IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation / Regulation of NF-kappa B signaling / Regulation of TBK1, IKKε (IKBKE)-mediated activation of IRF3, IRF7 / Regulation of pyruvate metabolism / Termination of translesion DNA synthesis / Ovarian tumor domain proteases / Negative regulators of DDX58/IFIH1 signaling / Negative regulation of FGFR1 signaling / Negative regulation of FGFR2 signaling / Negative regulation of FGFR3 signaling / Negative regulation of FGFR4 signaling / Negative regulation of MAPK pathway / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / Iron uptake and transport / Deactivation of the beta-catenin transactivating complex / Metalloprotease DUBs / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / Major pathway of rRNA processing in the nucleolus and cytosol / GTP hydrolysis and joining of the 60S ribosomal subunit / Activation of NF-kappaB in B cells / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / SCF(Skp2)-mediated degradation of p27/p21 / FCERI mediated NF-kB activation / Autodegradation of the E3 ubiquitin ligase COP1 / Asymmetric localization of PCP proteins / Degradation of AXIN / Degradation of DVL Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Jomaa A / Lee JH | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: Receptor compaction and GTPase rearrangement drive SRP-mediated cotranslational protein translocation into the ER. Authors: Jae Ho Lee / Ahmad Jomaa / SangYoon Chung / Yu-Hsien Hwang Fu / Ruilin Qian / Xuemeng Sun / Hao-Hsuan Hsieh / Sowmya Chandrasekar / Xiaotian Bi / Simone Mattei / Daniel Boehringer / Shimon ...Authors: Jae Ho Lee / Ahmad Jomaa / SangYoon Chung / Yu-Hsien Hwang Fu / Ruilin Qian / Xuemeng Sun / Hao-Hsuan Hsieh / Sowmya Chandrasekar / Xiaotian Bi / Simone Mattei / Daniel Boehringer / Shimon Weiss / Nenad Ban / Shu-Ou Shan /    Abstract: The conserved signal recognition particle (SRP) cotranslationally delivers ~30% of the proteome to the eukaryotic endoplasmic reticulum (ER). The molecular mechanism by which eukaryotic SRP ...The conserved signal recognition particle (SRP) cotranslationally delivers ~30% of the proteome to the eukaryotic endoplasmic reticulum (ER). The molecular mechanism by which eukaryotic SRP transitions from cargo recognition in the cytosol to protein translocation at the ER is not understood. Here, structural, biochemical, and single-molecule studies show that this transition requires multiple sequential conformational rearrangements in the targeting complex initiated by guanosine triphosphatase (GTPase)-driven compaction of the SRP receptor (SR). Disruption of these rearrangements, particularly in mutant SRP54 linked to severe congenital neutropenia, uncouples the SRP/SR GTPase cycle from protein translocation. Structures of targeting intermediates reveal the molecular basis of early SRP-SR recognition and emphasize the role of eukaryote-specific elements in regulating targeting. Our results provide a molecular model for the structural and functional transitions of SRP throughout the targeting cycle and show that these transitions provide important points for biological regulation that can be perturbed in genetic diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12303.map.gz emd_12303.map.gz | 321.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12303-v30.xml emd-12303-v30.xml emd-12303.xml emd-12303.xml | 80.8 KB 80.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_12303.png emd_12303.png | 85.5 KB | ||
| Filedesc metadata |  emd-12303.cif.gz emd-12303.cif.gz | 16.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12303 http://ftp.pdbj.org/pub/emdb/structures/EMD-12303 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12303 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12303 | HTTPS FTP |
-Validation report
| Summary document |  emd_12303_validation.pdf.gz emd_12303_validation.pdf.gz | 743 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12303_full_validation.pdf.gz emd_12303_full_validation.pdf.gz | 742.6 KB | Display | |
| Data in XML |  emd_12303_validation.xml.gz emd_12303_validation.xml.gz | 7.5 KB | Display | |
| Data in CIF |  emd_12303_validation.cif.gz emd_12303_validation.cif.gz | 8.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12303 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12303 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12303 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12303 | HTTPS FTP |
-Related structure data
| Related structure data |  7nfxMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12303.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12303.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | complex of RNC with SRPG226E and SRP receptor | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.062 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
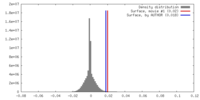
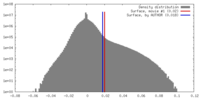
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : ribosome nascent chain complex with SRP and SRP receptor
+Supramolecule #1: ribosome nascent chain complex with SRP and SRP receptor
+Supramolecule #2: ribosomal proteins
+Supramolecule #3: SRP
+Supramolecule #4: Signal Sequence
+Macromolecule #1: SRP RNA 7SL
+Macromolecule #2: 28S ribosomal RNA
+Macromolecule #3: 5S ribosomal RNA
+Macromolecule #4: 5.8S ribosomal RNA
+Macromolecule #5: uL2
+Macromolecule #6: uL3
+Macromolecule #7: 60S ribosomal protein L4
+Macromolecule #8: 60S ribosomal protein L5
+Macromolecule #9: 60S ribosomal protein L6
+Macromolecule #10: uL30
+Macromolecule #11: 60S ribosomal protein L7a
+Macromolecule #12: 60S ribosomal protein L9
+Macromolecule #13: 60S ribosomal protein L10
+Macromolecule #14: Ribosomal protein L11
+Macromolecule #15: 60S ribosomal protein L13
+Macromolecule #16: 60S ribosomal protein L14
+Macromolecule #17: Ribosomal protein L15
+Macromolecule #18: uL13
+Macromolecule #19: uL22
+Macromolecule #20: eL18
+Macromolecule #21: 60S RIBOSOMAL PROTEIN EL19
+Macromolecule #22: eL20
+Macromolecule #23: eL21
+Macromolecule #24: Ribosomal protein L22
+Macromolecule #25: Ribosomal protein L23
+Macromolecule #26: Ribosomal protein L24
+Macromolecule #27: uL23
+Macromolecule #28: Ribosomal protein L26
+Macromolecule #29: 60S ribosomal protein L27
+Macromolecule #30: uL15
+Macromolecule #31: 60S ribosomal protein L29
+Macromolecule #32: eL30
+Macromolecule #33: eL31
+Macromolecule #34: eL32
+Macromolecule #35: eL33
+Macromolecule #36: 60S ribosomal protein L34
+Macromolecule #37: uL29
+Macromolecule #38: eL36
+Macromolecule #39: Ribosomal protein L37
+Macromolecule #40: eL38
+Macromolecule #41: eL39
+Macromolecule #42: 60S RIBOSOMAL PROTEIN EL40
+Macromolecule #43: 60s ribosomal protein l41
+Macromolecule #44: eL42
+Macromolecule #45: eL43
+Macromolecule #46: Signal recognition particle 19 kDa protein
+Macromolecule #47: eL28
+Macromolecule #48: Signal Sequence
+Macromolecule #49: Signal recognition particle 14 kDa protein
+Macromolecule #50: Signal recognition particle subunit SRP68
+Macromolecule #51: Signal recognition particle receptor subunit beta
+Macromolecule #52: Signal recognition particle 9 kDa protein
+Macromolecule #53: Signal recognition particle 54 kDa protein
+Macromolecule #54: Signal recognition particle receptor subunit alpha
+Macromolecule #55: Signal recognition particle subunit SRP72
+Macromolecule #56: ZINC ION
+Macromolecule #57: MAGNESIUM ION
+Macromolecule #58: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #59: PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. |
| Vitrification | Cryogen name: ETHANE-PROPANE / Instrument: FEI VITROBOT MARK IV |
| Details | in-vitro translation system |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL / In silico model: using SGD from particles |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 32881 |
| Initial angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)