[English] 日本語
 Yorodumi
Yorodumi- EMDB-12140: Cryo-EM structure of the outward open proton coupled folate trans... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12140 | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the outward open proton coupled folate transporter at pH 7.5 | |||||||||||||||||||||||||||||||||
 Map data Map data | globally sharpened map from cryosparc non uniform refinement | |||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||
 Keywords Keywords | transporter folate proton symporter / MEMBRANE PROTEIN | |||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationfolic acid:proton symporter activity / Metabolism of folate and pterines / Heme signaling / Iron uptake and transport / methotrexate transmembrane transporter activity / folate import across plasma membrane / folic acid binding / transmembrane transporter activity / transmembrane transport / basolateral plasma membrane ...folic acid:proton symporter activity / Metabolism of folate and pterines / Heme signaling / Iron uptake and transport / methotrexate transmembrane transporter activity / folate import across plasma membrane / folic acid binding / transmembrane transporter activity / transmembrane transport / basolateral plasma membrane / endosome / endosome membrane / apical plasma membrane / plasma membrane Similarity search - Function | |||||||||||||||||||||||||||||||||
| Biological species |   | |||||||||||||||||||||||||||||||||
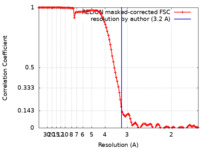
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||||||||||||||||||||||||||
 Authors Authors | Parker JL / Deme JC | |||||||||||||||||||||||||||||||||
| Funding support |  United Kingdom, 10 items United Kingdom, 10 items
| |||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Structural basis of antifolate recognition and transport by PCFT. Authors: Joanne L Parker / Justin C Deme / Gabriel Kuteyi / Zhiyi Wu / Jiandong Huo / I David Goldman / Raymond J Owens / Philip C Biggin / Susan M Lea / Simon Newstead /   Abstract: Folates (also known as vitamin B9) have a critical role in cellular metabolism as the starting point in the synthesis of nucleic acids, amino acids and the universal methylating agent S- ...Folates (also known as vitamin B9) have a critical role in cellular metabolism as the starting point in the synthesis of nucleic acids, amino acids and the universal methylating agent S-adenylsmethionine. Folate deficiency is associated with a number of developmental, immune and neurological disorders. Mammals cannot synthesize folates de novo; several systems have therefore evolved to take up folates from the diet and distribute them within the body. The proton-coupled folate transporter (PCFT) (also known as SLC46A1) mediates folate uptake across the intestinal brush border membrane and the choroid plexus, and is an important route for the delivery of antifolate drugs in cancer chemotherapy. How PCFT recognizes folates or antifolate agents is currently unclear. Here we present cryo-electron microscopy structures of PCFT in a substrate-free state and in complex with a new-generation antifolate drug (pemetrexed). Our results provide a structural basis for understanding antifolate recognition and provide insights into the pH-regulated mechanism of folate transport mediated by PCFT. | |||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12140.map.gz emd_12140.map.gz | 64.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12140-v30.xml emd-12140-v30.xml emd-12140.xml emd-12140.xml | 26 KB 26 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12140_fsc.xml emd_12140_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_12140.png emd_12140.png | 119.3 KB | ||
| Masks |  emd_12140_msk_1.map emd_12140_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12140.cif.gz emd-12140.cif.gz | 7.2 KB | ||
| Others |  emd_12140_additional_1.map.gz emd_12140_additional_1.map.gz emd_12140_half_map_1.map.gz emd_12140_half_map_1.map.gz emd_12140_half_map_2.map.gz emd_12140_half_map_2.map.gz | 62.6 MB 115.8 MB 115.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12140 http://ftp.pdbj.org/pub/emdb/structures/EMD-12140 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12140 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12140 | HTTPS FTP |
-Related structure data
| Related structure data |  7bc6MC  7bc7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12140.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12140.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | globally sharpened map from cryosparc non uniform refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
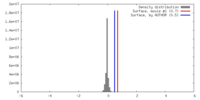
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12140_msk_1.map emd_12140_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
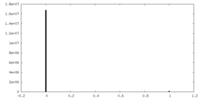
| Density Histograms |
-Additional map: unsharpened map from cryosparc non uniform refinement
| File | emd_12140_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map from cryosparc non uniform refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
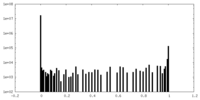
| Density Histograms |
-Half map: half map B from cryosparc non uniform refinement
| File | emd_12140_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B from cryosparc non uniform refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
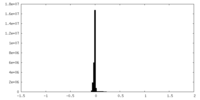
| Density Histograms |
-Half map: half map A from cryosparc non uniform refinement
| File | emd_12140_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A from cryosparc non uniform refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of the outward open proton coupled folate transporter wit...
| Entire | Name: Complex of the outward open proton coupled folate transporter with nanobody at pH 7.5 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of the outward open proton coupled folate transporter wit...
| Supramolecule | Name: Complex of the outward open proton coupled folate transporter with nanobody at pH 7.5 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: proton-coupled folate transporter
| Supramolecule | Name: proton-coupled folate transporter / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: nanobody
| Supramolecule | Name: nanobody / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Proton-coupled folate transporter
| Macromolecule | Name: Proton-coupled folate transporter / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 51.36657 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAPSDPPTA ATPPAPPPPA RRCLPAPSVE PLLFLATLAL GLQVPLATQY LWDRLGAERG YVGPNASSPH GCGNGSGAVD PLREEVEAL VAHWNLCINL GGFFVGLFSV TLFGPWSDSV GRRPVLVLPA VGMAVQAAVY LLVMYLRLHV AYLLLGRIIS G LLGDYNLI ...String: MAAPSDPPTA ATPPAPPPPA RRCLPAPSVE PLLFLATLAL GLQVPLATQY LWDRLGAERG YVGPNASSPH GCGNGSGAVD PLREEVEAL VAHWNLCINL GGFFVGLFSV TLFGPWSDSV GRRPVLVLPA VGMAVQAAVY LLVMYLRLHV AYLLLGRIIS G LLGDYNLI LAGCFASVAD SSNQRTRTFR VAILEACLGV AGMVASVGGG QWRKAEGYIN PFWLVLAASL AAALYAALCL QE TVKQRRA AKLLTLQHYK AVYKLYTAPE DLSSRRKLAL YSLAFFLLVT VHFGTKDLYV LYELGSPLCW ASDLIGYGSA ASY LAYLSS LGGLRLLQLC LEDTWVAEIG LISNIAGLVV ISLATTTPLM FTGYGIMFLS MAATPVIRAK LSKLVSETEQ GALF ASVAC VEGLCSLVAT GVFNSLYPST LHFMRGFPFL FGAILLLIPA AIMGWIEIQD SNLQYSHFSD ASSSPADGGE NLYFQ UniProtKB: Proton-coupled folate transporter |
-Macromolecule #2: nanobody
| Macromolecule | Name: nanobody / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.460006 KDa |
| Sequence | String: GPSQVQLVES GGGLVQPGGS LRLSCAASGF TFSRYWMYWV RQAPGKGPEW LSHMNPSGSD IKYTDSVKGR FTISRDNAKN TLYLQMNSL KPDDTAVYYC VADRRALGSP EYWGQGTQVT VSSA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 59.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)