+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1048 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Three-dimensional structure of bacteriophage T4 baseplate. | |||||||||
 Map data Map data | map contains the baseplate and the proximal part of the tail tube | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont entry into host cell via disruption of host cell wall peptidoglycan / virus tail, baseplate / viral tail assembly / virus tail, fiber / symbiont entry into host cell via disruption of host cell envelope / virus tail / symbiont entry into host / viral release from host cell / peptidoglycan catabolic process / cell wall macromolecule catabolic process ...symbiont entry into host cell via disruption of host cell wall peptidoglycan / virus tail, baseplate / viral tail assembly / virus tail, fiber / symbiont entry into host cell via disruption of host cell envelope / virus tail / symbiont entry into host / viral release from host cell / peptidoglycan catabolic process / cell wall macromolecule catabolic process / lysozyme / lysozyme activity / killing of cells of another organism / entry receptor-mediated virion attachment to host cell / defense response to bacterium / symbiont entry into host cell / metal ion binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 12.0 Å | |||||||||
 Authors Authors | Kostyuchenko VA / Leiman PG / Chipman PR / Kanamaru S / vanRaaij MJ / Arisaka F / Mesyanzhinov VV / Rossmann MG | |||||||||
 Citation Citation |  Journal: Nat Struct Biol / Year: 2003 Journal: Nat Struct Biol / Year: 2003Title: Three-dimensional structure of bacteriophage T4 baseplate. Authors: Victor A Kostyuchenko / Petr G Leiman / Paul R Chipman / Shuji Kanamaru / Mark J van Raaij / Fumio Arisaka / Vadim V Mesyanzhinov / Michael G Rossmann /  Abstract: The baseplate of bacteriophage T4 is a multiprotein molecular machine that controls host cell recognition, attachment, tail sheath contraction and viral DNA ejection. We report here the three- ...The baseplate of bacteriophage T4 is a multiprotein molecular machine that controls host cell recognition, attachment, tail sheath contraction and viral DNA ejection. We report here the three-dimensional structure of the baseplate-tail tube complex determined to a resolution of 12 A by cryoelectron microscopy. The baseplate has a six-fold symmetric, dome-like structure approximately 520 A in diameter and approximately 270 A long, assembled around a central hub. A 940 A-long and 96 A-diameter tail tube, coaxial with the hub, is connected to the top of the baseplate. At the center of the dome is a needle-like structure that was previously identified as a cell puncturing device. We have identified the locations of six proteins with known atomic structures, and established the position and shape of several other baseplate proteins. The baseplate structure suggests a mechanism of baseplate triggering and structural transition during the initial stages of T4 infection. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1048.map.gz emd_1048.map.gz | 3.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1048-v30.xml emd-1048-v30.xml emd-1048.xml emd-1048.xml | 14 KB 14 KB | Display Display |  EMDB header EMDB header |
| Images |  1048.gif 1048.gif | 53.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1048 http://ftp.pdbj.org/pub/emdb/structures/EMD-1048 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1048 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1048 | HTTPS FTP |
-Related structure data
| Related structure data |  1pdfMC  1pdiMC  1pdjMC  1pdlMC  1pdmMC  1pdpMC  2fl8M  3h3wM M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1048.map.gz / Format: CCP4 / Size: 28.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1048.map.gz / Format: CCP4 / Size: 28.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map contains the baseplate and the proximal part of the tail tube | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.97872 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
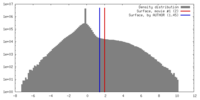
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriophage T4 baseplate-tail tube assembly
| Entire | Name: Bacteriophage T4 baseplate-tail tube assembly |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriophage T4 baseplate-tail tube assembly
| Supramolecule | Name: Bacteriophage T4 baseplate-tail tube assembly / type: sample / ID: 1000 / Oligomeric state: the complex has sixfold symmetry / Number unique components: 1 |
|---|
-Macromolecule #1: baseplate-tail tube complex
| Macromolecule | Name: baseplate-tail tube complex / type: protein_or_peptide / ID: 1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Enterobacteria phage T4 (virus) / Strain: T4 18-,23- / synonym: bacteriophage T4 / Organelle: baseplate and tail tube Enterobacteria phage T4 (virus) / Strain: T4 18-,23- / synonym: bacteriophage T4 / Organelle: baseplate and tail tube |
| Recombinant expression | Organism: Escherichia coli BE |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




















 Processing
Processing Electron microscopy
Electron microscopy