[English] 日本語
 Yorodumi
Yorodumi- EMDB-0931: Cryo-EM structure of amyloid fibril formed by full-length human p... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0931 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of amyloid fibril formed by full-length human prion protein | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Amyloid fibril / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of amyloid precursor protein catabolic process / regulation of glutamate receptor signaling pathway / lamin binding / aspartic-type endopeptidase inhibitor activity / regulation of calcium ion import across plasma membrane / glycosaminoglycan binding / NCAM1 interactions / type 5 metabotropic glutamate receptor binding / positive regulation of glutamate receptor signaling pathway / negative regulation of interleukin-17 production ...negative regulation of amyloid precursor protein catabolic process / regulation of glutamate receptor signaling pathway / lamin binding / aspartic-type endopeptidase inhibitor activity / regulation of calcium ion import across plasma membrane / glycosaminoglycan binding / NCAM1 interactions / type 5 metabotropic glutamate receptor binding / positive regulation of glutamate receptor signaling pathway / negative regulation of interleukin-17 production / cupric ion binding / ATP-dependent protein binding / regulation of potassium ion transmembrane transport / negative regulation of protein processing / negative regulation of dendritic spine maintenance / dendritic spine maintenance / extrinsic component of membrane / negative regulation of calcineurin-NFAT signaling cascade / negative regulation of interleukin-2 production / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / negative regulation of activated T cell proliferation / negative regulation of amyloid-beta formation / response to amyloid-beta / negative regulation of type II interferon production / cuprous ion binding / negative regulation of long-term synaptic potentiation / intracellular copper ion homeostasis / negative regulation of T cell receptor signaling pathway / positive regulation of protein targeting to membrane / response to cadmium ion / long-term memory / neuron projection maintenance / inclusion body / positive regulation of calcium-mediated signaling / cellular response to copper ion / molecular function activator activity / positive regulation of protein localization to plasma membrane / molecular condensate scaffold activity / tubulin binding / protein homooligomerization / protein destabilization / cellular response to xenobiotic stimulus / cellular response to amyloid-beta / terminal bouton / positive regulation of neuron apoptotic process / amyloid-beta binding / protein-folding chaperone binding / signaling receptor activity / protease binding / response to oxidative stress / nuclear membrane / microtubule binding / molecular adaptor activity / transmembrane transporter binding / learning or memory / regulation of cell cycle / postsynaptic density / intracellular signal transduction / postsynapse / membrane raft / copper ion binding / external side of plasma membrane / dendrite / negative regulation of apoptotic process / protein-containing complex binding / cell surface / negative regulation of transcription by RNA polymerase II / endoplasmic reticulum / Golgi apparatus / extracellular exosome / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
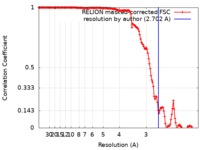
| Method | helical reconstruction / cryo EM / Resolution: 2.702 Å | |||||||||
 Authors Authors | Wang LQ / Zhao K | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2020 Journal: Nat Struct Mol Biol / Year: 2020Title: Cryo-EM structure of an amyloid fibril formed by full-length human prion protein. Authors: Li-Qiang Wang / Kun Zhao / Han-Ye Yuan / Qiang Wang / Zeyuan Guan / Jing Tao / Xiang-Ning Li / Yunpeng Sun / Chuan-Wei Yi / Jie Chen / Dan Li / Delin Zhang / Ping Yin / Cong Liu / Yi Liang /  Abstract: Prion diseases are caused by the misfolding of prion protein (PrP). Misfolded PrP forms protease-resistant aggregates in vivo (PrP) that are able to template the conversion of the native form of the ...Prion diseases are caused by the misfolding of prion protein (PrP). Misfolded PrP forms protease-resistant aggregates in vivo (PrP) that are able to template the conversion of the native form of the protein (PrP), a property shared by in vitro-produced PrP fibrils. Here we produced amyloid fibrils in vitro from recombinant, full-length human PrP (residues 23-231) and determined their structure using cryo-EM, building a model for the fibril core comprising residues 170-229. The PrP fibril consists of two protofibrils intertwined in a left-handed helix. Lys194 and Glu196 from opposing subunits form salt bridges, creating a hydrophilic cavity at the interface of the two protofibrils. By comparison with the structure of PrP, we propose that two α-helices in the C-terminal domain of PrP are converted into β-strands stabilized by a disulfide bond in the PrP fibril. Our data suggest that different PrP mutations may play distinct roles in modulating the conformational conversion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0931.map.gz emd_0931.map.gz | 18.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0931-v30.xml emd-0931-v30.xml emd-0931.xml emd-0931.xml | 9.2 KB 9.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0931_fsc.xml emd_0931_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_0931.png emd_0931.png | 35.9 KB | ||
| Filedesc metadata |  emd-0931.cif.gz emd-0931.cif.gz | 4.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0931 http://ftp.pdbj.org/pub/emdb/structures/EMD-0931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0931 | HTTPS FTP |
-Related structure data
| Related structure data |  6lniMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0931.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0931.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.014 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
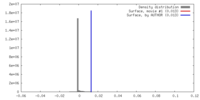
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human prion protein amyloid fibril
| Entire | Name: human prion protein amyloid fibril |
|---|---|
| Components |
|
-Supramolecule #1: human prion protein amyloid fibril
| Supramolecule | Name: human prion protein amyloid fibril / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Major prion protein
| Macromolecule | Name: Major prion protein / type: protein_or_peptide / ID: 1 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.996355 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKRPKPGGW NTGGSRYPGQ GSPGGNRYPP QGGGGWGQPH GGGWGQPHGG GWGQPHGGGW GQPHGGGWGQ GGGTHSQWNK PSKPKTNMK HMAGAAAAGA VVGGLGGYML GSAMSRPIIH FGSDYEDRYY RENMHRYPNQ VYYRPMDEYS NQNNFVHDCV N ITIKQHTV ...String: MKKRPKPGGW NTGGSRYPGQ GSPGGNRYPP QGGGGWGQPH GGGWGQPHGG GWGQPHGGGW GQPHGGGWGQ GGGTHSQWNK PSKPKTNMK HMAGAAAAGA VVGGLGGYML GSAMSRPIIH FGSDYEDRYY RENMHRYPNQ VYYRPMDEYS NQNNFVHDCV N ITIKQHTV TTTTKGENFT ETDVKMMERV VEQMCITQYE RESQAYYQRG SS UniProtKB: Major prion protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 64.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)