+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0915 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | WT transporter state1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | calcium / METAL TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationlongitudinal sarcoplasmic reticulum / ER-nucleus signaling pathway / positive regulation of endoplasmic reticulum calcium ion concentration / P-type calcium transporter activity involved in regulation of cardiac muscle cell membrane potential / calcium ion transport from cytosol to endoplasmic reticulum / regulation of calcium ion-dependent exocytosis of neurotransmitter / calcium ion-transporting ATPase complex / T-tubule organization / regulation of cardiac muscle cell action potential involved in regulation of contraction / regulation of cardiac muscle cell membrane potential ...longitudinal sarcoplasmic reticulum / ER-nucleus signaling pathway / positive regulation of endoplasmic reticulum calcium ion concentration / P-type calcium transporter activity involved in regulation of cardiac muscle cell membrane potential / calcium ion transport from cytosol to endoplasmic reticulum / regulation of calcium ion-dependent exocytosis of neurotransmitter / calcium ion-transporting ATPase complex / T-tubule organization / regulation of cardiac muscle cell action potential involved in regulation of contraction / regulation of cardiac muscle cell membrane potential / platelet dense tubular network membrane / calcium ion import into sarcoplasmic reticulum / Pre-NOTCH Processing in Golgi / sarcoplasmic reticulum calcium ion transport / ribbon synapse / P-type Ca2+ transporter / negative regulation of heart contraction / P-type calcium transporter activity / regulation of the force of heart contraction / transition between fast and slow fiber / endoplasmic reticulum calcium ion homeostasis / S100 protein binding / cardiac muscle hypertrophy in response to stress / relaxation of cardiac muscle / regulation of cardiac muscle contraction by calcium ion signaling / Reduction of cytosolic Ca++ levels / organelle localization by membrane tethering / mitochondrion-endoplasmic reticulum membrane tethering / autophagosome membrane docking / lncRNA binding / Ion transport by P-type ATPases / epidermis development / regulation of cardiac conduction / positive regulation of heart rate / autophagosome assembly / Ion homeostasis / sarcoplasmic reticulum membrane / positive regulation of cardiac muscle cell apoptotic process / response to endoplasmic reticulum stress / sarcoplasmic reticulum / calcium channel regulator activity / neuron cellular homeostasis / calcium ion transmembrane transport / intracellular calcium ion homeostasis / cellular response to oxidative stress / monoatomic ion transmembrane transport / transmembrane transporter binding / cell adhesion / calcium ion binding / endoplasmic reticulum membrane / enzyme binding / endoplasmic reticulum / ATP hydrolysis activity / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Zhang Y / Tsutsumi A | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2020 Journal: Sci Adv / Year: 2020Title: Cryo-EM structures of SERCA2b reveal the mechanism of regulation by the luminal extension tail. Authors: Yuxia Zhang / Michio Inoue / Akihisa Tsutsumi / Satoshi Watanabe / Tomohiro Nishizawa / Kazuhiro Nagata / Masahide Kikkawa / Kenji Inaba /  Abstract: Sarco/endoplasmic reticulum Ca ATPase (SERCA) pumps Ca from the cytosol into the ER and maintains the cellular calcium homeostasis. Herein, we present cryo-electron microscopy (cryo-EM) structures of ...Sarco/endoplasmic reticulum Ca ATPase (SERCA) pumps Ca from the cytosol into the ER and maintains the cellular calcium homeostasis. Herein, we present cryo-electron microscopy (cryo-EM) structures of human SERCA2b in E1∙2Ca-adenylyl methylenediphosphonate (AMPPCP) and E2-BeF states at 2.9- and 2.8-Å resolutions, respectively. The structures revealed that the luminal extension tail (LE) characteristic of SERCA2b runs parallel to the lipid-water boundary near the luminal ends of transmembrane (TM) helices TM10 and TM7 and approaches the luminal loop flanked by TM7 and TM8. While the LE served to stabilize the cytosolic and TM domain arrangement of SERCA2b, deletion of the LE rendered the overall conformation resemble that of SERCA1a and SERCA2a and allowed multiple conformations. Thus, the LE appears to play a critical role in conformational regulation in SERCA2b, which likely explains the different kinetic properties of SERCA2b from those of other isoforms lacking the LE. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0915.map.gz emd_0915.map.gz | 2.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0915-v30.xml emd-0915-v30.xml emd-0915.xml emd-0915.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0915_fsc.xml emd_0915_fsc.xml | 6 KB | Display |  FSC data file FSC data file |
| Images |  emd_0915.png emd_0915.png | 47.1 KB | ||
| Masks |  emd_0915_msk_1.map emd_0915_msk_1.map | 18.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-0915.cif.gz emd-0915.cif.gz | 6.8 KB | ||
| Others |  emd_0915_additional.map.gz emd_0915_additional.map.gz emd_0915_additional_1.map.gz emd_0915_additional_1.map.gz emd_0915_half_map_1.map.gz emd_0915_half_map_1.map.gz emd_0915_half_map_2.map.gz emd_0915_half_map_2.map.gz | 13.8 MB 13.8 MB 13.8 MB 13.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0915 http://ftp.pdbj.org/pub/emdb/structures/EMD-0915 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0915 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0915 | HTTPS FTP |
-Related structure data
| Related structure data |  6llyMC  0912C  0924C  0925C  0926C  0927C  0928C  6lleC  6ln5C  6ln6C  6ln7C  6ln8C  6ln9C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0915.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0915.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.245 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
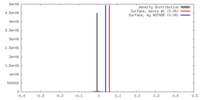
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0915_msk_1.map emd_0915_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: no sharpen map
| File | emd_0915_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | no sharpen map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: no sharpen map
| File | emd_0915_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | no sharpen map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: unfiltered half1 map
| File | emd_0915_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered half1 map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: unfiltered half2 map
| File | emd_0915_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered half2 map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SERCA2b with BeF3
| Entire | Name: SERCA2b with BeF3 |
|---|---|
| Components |
|
-Supramolecule #1: SERCA2b with BeF3
| Supramolecule | Name: SERCA2b with BeF3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 110 kDa/nm |
-Macromolecule #1: Sarcoplasmic/endoplasmic reticulum calcium ATPase 2
| Macromolecule | Name: Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: P-type Ca2+ transporter |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 117.764875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGGVAMPGAE DDVVRENLYF QGKDGLAAME NAHTKTVEEV LGHFGVNEST GLSLEQVKKL KERWGSNELP AEEGKTLLEL VIEQFEDLL VRILLLAACI SFVLAWFEEG EETITAFVEP FVILLILVAN AIVGVWQERN AENAIEALKE YEPEMGKVYR Q DRKSVQRI ...String: MGGVAMPGAE DDVVRENLYF QGKDGLAAME NAHTKTVEEV LGHFGVNEST GLSLEQVKKL KERWGSNELP AEEGKTLLEL VIEQFEDLL VRILLLAACI SFVLAWFEEG EETITAFVEP FVILLILVAN AIVGVWQERN AENAIEALKE YEPEMGKVYR Q DRKSVQRI KAKDIVPGDI VEIAVGDKVP ADIRLTSIKS TTLRVDQSIL TGESVSVIKH TDPVPDPRAV NQDKKNMLFS GT NIAAGKA MGVVVATGVN TEIGKIRDEM VATEQERTPL QQKLDEFGEQ LSKVISLICI AVWIINIGHF NDPVHGGSWI RGA IYYFKI AVALAVAAIP EGLPAVITTC LALGTRRMAK KNAIVRSLPS VETLGCTSVI CSDKTGTLTT NQMSVCRMFI LDRV EGDTC SLNEFTITGS TYAPIGEVHK DDKPVNCHQY DGLVELATIC ALCNDSALDY NEAKGVYEKV GEATETALTC LVEKM NVFD TELKGLSKIE RANACNSVIK QLMKKEFTLE FSRDRKSMSV YCTPNKPSRT SMSKMFVKGA PEGVIDRCTH IRVGST KVP MTSGVKQKIM SVIREWGSGS DTLRCLALAT HDNPLRREEM HLEDSANFIK YETNLTFVGC VGMLDPPRIE VASSVKL CR QAGIRVIMIT GDNKGTAVAI CRRIGIFGQD EDVTSKAFTG REFDELNPSA QRDACLNARC FARVEPSHKS KIVEFLQS F DEITAMTGDG VNDAPALKKA EIGIAMGSGT AVAKTASEMV LADDNFSTIV AAVEEGRAIY NNMKQFIRYL ISSNVGEVV CIFLTAALGF PEALIPVQLL WVNLVTDGLP ATALGFNPPD LDIMNKPPRN PKEPLISGWL FFRYLAIGCY VGAATVGAAA WWFIAADGG PRVSFYQLSH FLQCKEDNPD FEGVDCAIFE SPYPMTMALS VLVTIEMCNA LNSLSENQSL LRMPPWENIW L VGSICLSM SLHFLILYVE PLPLIFQITP LNVTQWLMVL KISLPVILMD ETLKFVARNY LEPGKECVQP ATKSCSFSAC TD GISWPFV LLIMPLVIWV YSTDTNFSDM FWS UniProtKB: Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 |
-Macromolecule #2: BERYLLIUM TRIFLUORIDE ION
| Macromolecule | Name: BERYLLIUM TRIFLUORIDE ION / type: ligand / ID: 2 / Number of copies: 1 / Formula: BEF |
|---|---|
| Molecular weight | Theoretical: 66.007 Da |
| Chemical component information |  ChemComp-BEF: |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)