+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0623 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the AAV2 with its Cell Receptor, AAVR | |||||||||
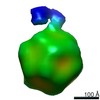
 Map data Map data | AAV2 bound with AAVR(class 2) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Adeno-associated virus Adeno-associated virus | |||||||||
| Method | electron tomography / cryo EM / Resolution: 20.0 Å | |||||||||
 Authors Authors | Hu GQ / Meyer NL / Stagg SM / Chapman MS / Davulcu O / Xie Q / Noble AJ / Yoshioka C / Gingerich D / Trzynka A / David L | |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: Structure of the gene therapy vector, adeno-associated virus with its cell receptor, AAVR. Authors: Nancy L Meyer / Guiqing Hu / Omar Davulcu / Qing Xie / Alex J Noble / Craig Yoshioka / Drew S Gingerich / Andrew Trzynka / Larry David / Scott M Stagg / Michael Stewart Chapman /  Abstract: Adeno-associated virus (AAV) vectors are preeminent in emerging clinical gene therapies. Generalizing beyond the most tractable genetic diseases will require modulation of cell specificity and immune ...Adeno-associated virus (AAV) vectors are preeminent in emerging clinical gene therapies. Generalizing beyond the most tractable genetic diseases will require modulation of cell specificity and immune neutralization. Interactions of AAV with its cellular receptor, AAVR, are key to understanding cell-entry and trafficking with the rigor needed to engineer tissue-specific vectors. -electron tomography shows ordered binding of part of the flexible receptor to the viral surface, with distal domains in multiple conformations. Regions of the virus and receptor in close physical proximity can be identified by cross-linking/mass spectrometry. -electron microscopy with a two-domain receptor fragment reveals the interactions at 2.4 Å resolution. AAVR binds between AAV's spikes on a plateau that is conserved, except in one clade whose structure is AAVR-incompatible. AAVR's footprint overlaps the epitopes of several neutralizing antibodies, prompting a re-evaluation of neutralization mechanisms. The structure provides a roadmap for experimental probing and manipulation of viral-receptor interactions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0623.map.gz emd_0623.map.gz | 132.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0623-v30.xml emd-0623-v30.xml emd-0623.xml emd-0623.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0623.png emd_0623.png | 52.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0623 http://ftp.pdbj.org/pub/emdb/structures/EMD-0623 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0623 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0623 | HTTPS FTP |
-Validation report
| Summary document |  emd_0623_validation.pdf.gz emd_0623_validation.pdf.gz | 79.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0623_full_validation.pdf.gz emd_0623_full_validation.pdf.gz | 78.7 KB | Display | |
| Data in XML |  emd_0623_validation.xml.gz emd_0623_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0623 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0623 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0623 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0623 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0623.map.gz / Format: CCP4 / Size: 182.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0623.map.gz / Format: CCP4 / Size: 182.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AAV2 bound with AAVR(class 2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 15.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : AAV2 complex with AAVR receptor
| Entire | Name: AAV2 complex with AAVR receptor |
|---|---|
| Components |
|
-Supramolecule #1: AAV2 complex with AAVR receptor
| Supramolecule | Name: AAV2 complex with AAVR receptor / type: complex / ID: 1 / Parent: 0 / Details: class 1 |
|---|---|
| Source (natural) | Organism:   Adeno-associated virus / Strain: hybrid of serotypes 2, 8, and 9 Adeno-associated virus / Strain: hybrid of serotypes 2, 8, and 9 |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 3.75 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: blot force = 1, blot time = 3 seconds, total blots = 1 | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 4 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Sectioning | Other: NO SECTIONING |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 290.0 K / Max: 300.0 K |
| Details | preliminary grid screening was performed manually |
| Image recording | Film or detector model: DIRECT ELECTRON DE-20 (5k x 3k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 5000 pixel / Digitization - Dimensions - Height: 3000 pixel / Digitization - Sampling interval: 10.0 µm / Digitization - Frames/image: 1-7 / Number grids imaged: 1 / Average exposure time: 1.5 sec. / Average electron dose: 1.42 e/Å2 / Details: none |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 0.011 µm / Calibrated defocus min: 9.0 µm / Calibrated magnification: 18000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 0.011 µm / Nominal defocus min: 9.0 µm / Nominal magnification: 18000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | nond |
|---|---|
| Final reconstruction | Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: OTHER / Software - Name: Dynamo (ver. 1.0) / Software - details: none / Details: none / Number images used: 280 |
| CTF correction | Software - Name: TOMOCTF (ver. 1.0) / Software - details: none / Details: tomoctf |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: correlation coeeficient |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















