[English] 日本語
 Yorodumi
Yorodumi- EMDB-0401: MERS-CoV complex with human neutralizing LCA60 antibody Fab fragm... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0401 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | MERS-CoV complex with human neutralizing LCA60 antibody Fab fragment (state 1) | |||||||||
 Map data Map data | MERS-CoV complex with human neutralizing LCA60 antibody Fab fragment (state 1), primary map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | coronavirus spike glycoprotein / MERS-CoV / SARS-CoV / human neutralizing antibodies / Structural Genomics / Seattle Structural Genomics Center for Infectious Disease / SSGCID / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane / membrane Similarity search - Function | |||||||||
| Biological species |    Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Walls AC / Xiong X | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2019 Journal: Cell / Year: 2019Title: Unexpected Receptor Functional Mimicry Elucidates Activation of Coronavirus Fusion. Authors: Alexandra C Walls / Xiaoli Xiong / Young-Jun Park / M Alejandra Tortorici / Joost Snijder / Joel Quispe / Elisabetta Cameroni / Robin Gopal / Mian Dai / Antonio Lanzavecchia / Maria Zambon / ...Authors: Alexandra C Walls / Xiaoli Xiong / Young-Jun Park / M Alejandra Tortorici / Joost Snijder / Joel Quispe / Elisabetta Cameroni / Robin Gopal / Mian Dai / Antonio Lanzavecchia / Maria Zambon / Félix A Rey / Davide Corti / David Veesler /     Abstract: Recent outbreaks of severe acute respiratory syndrome and Middle East respiratory syndrome, along with the threat of a future coronavirus-mediated pandemic, underscore the importance of finding ways ...Recent outbreaks of severe acute respiratory syndrome and Middle East respiratory syndrome, along with the threat of a future coronavirus-mediated pandemic, underscore the importance of finding ways to combat these viruses. The trimeric spike transmembrane glycoprotein S mediates entry into host cells and is the major target of neutralizing antibodies. To understand the humoral immune response elicited upon natural infections with coronaviruses, we structurally characterized the SARS-CoV and MERS-CoV S glycoproteins in complex with neutralizing antibodies isolated from human survivors. Although the two antibodies studied blocked attachment to the host cell receptor, only the anti-SARS-CoV S antibody triggered fusogenic conformational changes via receptor functional mimicry. These results provide a structural framework for understanding coronavirus neutralization by human antibodies and shed light on activation of coronavirus membrane fusion, which takes place through a receptor-driven ratcheting mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0401.map.gz emd_0401.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0401-v30.xml emd-0401-v30.xml emd-0401.xml emd-0401.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0401.png emd_0401.png | 77.9 KB | ||
| Filedesc metadata |  emd-0401.cif.gz emd-0401.cif.gz | 7.2 KB | ||
| Others |  emd_0401_additional.map.gz emd_0401_additional.map.gz | 108.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0401 http://ftp.pdbj.org/pub/emdb/structures/EMD-0401 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0401 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0401 | HTTPS FTP |
-Related structure data
| Related structure data |  6nb3MC  0402C  0403C  0404C  6nb4C  6nb5C  6nb6C  6nb7C  6nb8C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0401.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0401.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | MERS-CoV complex with human neutralizing LCA60 antibody Fab fragment (state 1), primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
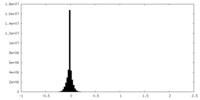
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: MERS-CoV complex with human neutralizing LCA60 antibody Fab...
| File | emd_0401_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | MERS-CoV complex with human neutralizing LCA60 antibody Fab fragment (state 1), Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
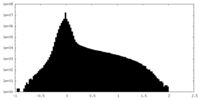
| Density Histograms |
- Sample components
Sample components
-Entire : MERS-CoV complex with human neutralizing LCA60 antibody Fab fragm...
| Entire | Name: MERS-CoV complex with human neutralizing LCA60 antibody Fab fragment (state 1) |
|---|---|
| Components |
|
-Supramolecule #1: MERS-CoV complex with human neutralizing LCA60 antibody Fab fragm...
| Supramolecule | Name: MERS-CoV complex with human neutralizing LCA60 antibody Fab fragment (state 1) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 572 MDa |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 149.172062 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GTVDVGPDSV KSACIEVDIQ QTFFDKTWPR PIDVSKADGI IYPQGRTYSN ITITYQGLF PYQGDHGDMY VYSAGHATGT TPQKLFVANY SQDVKQFANG FVVRIGAAAN STGTVIISPS TSATIRKIYP A FMLGSSVG ...String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GTVDVGPDSV KSACIEVDIQ QTFFDKTWPR PIDVSKADGI IYPQGRTYSN ITITYQGLF PYQGDHGDMY VYSAGHATGT TPQKLFVANY SQDVKQFANG FVVRIGAAAN STGTVIISPS TSATIRKIYP A FMLGSSVG NFSDGKMGRF FNHTLVLLPD GCGTLLRAFY CILEPRSGNH CPAGNSYTSF ATYHTPATDC SDGNYNRNAS LN SFKEYFN LRNCTFMYTY NITEDEILEW FGITQTAQGV HLFSSRYVDL YGGNMFQFAT LPVYDTIKYY SIIPHSIRSI QSD RKAWAA FYVYKLQPLT FLLDFSVDGY IRRAIDCGFN DLSQLHCSYE SFDVESGVYS VSSFEAKPSG SVVEQAEGVE CDFS PLLSG TPPQVYNFKR LVFTNCNYNL TKLLSLFSVN DFTCSQISPA AIASNCYSSL ILDYFSYPLS MKSDLSVSSA GPISQ FNYK QSFSNPTCLI LATVPHNLTT ITKPLKYSYI NKCSRLLSDD RTEVPQLVNA NQYSPCVSIV PSTVWEDGDY YRKQLS PLE GGGWLVASGS TVAMTEQLQM GFGITVQYGT DTNSVCPKLE FANDTKIASQ LGNCVEYSLY GVSGRGVFQN CTAVGVR QQ RFVYDAYQNL VGYYSDDGNY YCLRACVSVP VSVIYDKETK THATLFGSVA CEHISSTMSQ YSRSTRSMLK RRDSTYGP L QTPVGCVLGL VNSSLFVEDC KLPLGQSLCA LPDTPSTLTP ASVGSVPGEM RLASIAFNHP IQVDQLNSSY FKLSIPTNF SFGVTQEYIQ TTIQKVTVDC KQYVCNGFQK CEQLLREYGQ FCSKINQALH GANLRQDDSV RNLFASVKSS QSSPIIPGFG GDFNLTLLE PVSISTGSRS ARSAIEDLLF DKVTIADPGY MQGYDDCMQQ GPASARDLIC AQYVAGYKVL PPLMDVNMEA A YTSSLLGS IAGVGWTAGL SSFAAIPFAQ SIFYRLNGVG ITQQVLSENQ KLIANKFNQA LGAMQTGFTT TNEAFQKVQD AV NNNAQAL SKLASELSNT FGAISASIGD IIQRLDPPEQ DAQIDRLING RLTTLNAFVA QQLVRSESAA LSAQLAKDKV NEC VKAQSK RSGFCGQGTH IVSFVVNAPN GLYFMHVGYY PSNHIEVVSA YGLCDAANPT NCIAPVNGYF IKTNNTRIVD EWSY TGSSF YAPEPITSLN TKYVAPQVTY QNISTNLPPP LLGNSTGIDF QDELDEFFKN VSTSIPNFGS LTQINTTLLD LTYEM LSLQ QVVKALNESY IDLKELGNYT YYNKGSGREN LYFQGGGGSG YIPEAPRDGQ AYVRKDGEWV LLSTFLGHHH HHHHH UniProtKB: Spike glycoprotein |
-Macromolecule #2: LCA60 light chain
| Macromolecule | Name: LCA60 light chain / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.25939 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSALTQPASV SGSPGQSITI SCTGTSSDVG TYDLVSWYQQ HPGKSPKLMI YADIKRPSGV SHRFSGSKSG NTASLTISGL QSADEADYY CCLYAGSSTS VIFGGGTKVT |
-Macromolecule #3: LCA60 heavy chain
| Macromolecule | Name: LCA60 heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.920568 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLLESGGG LVKPGGSLRL SCEASGLTFS NVWMSWVRQA PGKGLEWVGR IKRKSEGATT DYGAPVKGRF TLSRDDSKNT VYLQMNSLK IDDTAVYYCS TLTRGGDVWS SSYYFDYWGQ GALVTVSS |
-Macromolecule #10: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 10 / Number of copies: 5 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: PDB 5W9J |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 54071 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION (ver. 3.0) |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-6nb3: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)