[English] 日本語
 Yorodumi
Yorodumi- EMDB-0343: MicroED structure of Proteinase K at 2.75A resolution from a sing... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0343 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | MicroED structure of Proteinase K at 2.75A resolution from a single milled crystal. | |||||||||
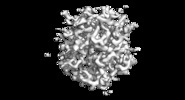
 Map data Map data | MicroED 2Fo-Fc map of proteinase K at 2.75A resolution from a single milled crystal. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | broad-spectrum serum proteinase / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationpeptidase K / serine-type endopeptidase activity / proteolysis / extracellular region / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Engyodontium album (fungus) Engyodontium album (fungus) | |||||||||
| Method | electron crystallography / cryo EM / Resolution: 2.75 Å | |||||||||
 Authors Authors | Martynowycz MW / Zhao W | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2019 Journal: Structure / Year: 2019Title: Collection of Continuous Rotation MicroED Data from Ion Beam-Milled Crystals of Any Size. Authors: Michael W Martynowycz / Wei Zhao / Johan Hattne / Grant J Jensen / Tamir Gonen /  Abstract: Microcrystal electron diffraction (MicroED) allows for macromolecular structure solution from nanocrystals. To create crystals of suitable size for MicroED data collection, sample preparation ...Microcrystal electron diffraction (MicroED) allows for macromolecular structure solution from nanocrystals. To create crystals of suitable size for MicroED data collection, sample preparation typically involves sonication or pipetting a slurry of crystals from a crystallization drop. The resultant crystal fragments are fragile and the quality of the data that can be obtained from them is sensitive to subsequent sample preparation for cryoelectron microscopy as interactions in the water-air interface can damage crystals during blotting. Here, we demonstrate the use of a focused ion beam to generate lamellae of macromolecular protein crystals for continuous rotation MicroED that are of ideal thickness, easy to locate, and require no blotting optimization. In this manner, crystals of nearly any size may be scooped and milled to desired dimensions prior to data collection, thus streamlining the methodology for sample preparation for MicroED. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0343.map.gz emd_0343.map.gz | 1.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0343-v30.xml emd-0343-v30.xml emd-0343.xml emd-0343.xml | 14.7 KB 14.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0343.png emd_0343.png | 94.9 KB | ||
| Filedesc metadata |  emd-0343.cif.gz emd-0343.cif.gz | 6.2 KB | ||
| Filedesc structureFactors |  emd_0343_sf.cif.gz emd_0343_sf.cif.gz | 316.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0343 http://ftp.pdbj.org/pub/emdb/structures/EMD-0343 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0343 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0343 | HTTPS FTP |
-Related structure data
| Related structure data |  6n4uMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0343.map.gz / Format: CCP4 / Size: 1.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0343.map.gz / Format: CCP4 / Size: 1.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | MicroED 2Fo-Fc map of proteinase K at 2.75A resolution from a single milled crystal. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X: 0.6746 Å / Y: 0.6746 Å / Z: 0.66669 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 96 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Proteinase K
| Entire | Name: Proteinase K |
|---|---|
| Components |
|
-Supramolecule #1: Proteinase K
| Supramolecule | Name: Proteinase K / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Engyodontium album (fungus) Engyodontium album (fungus) |
| Molecular weight | Theoretical: 28.93 KDa |
-Macromolecule #1: Proteinase K
| Macromolecule | Name: Proteinase K / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: peptidase K |
|---|---|
| Source (natural) | Organism:  Engyodontium album (fungus) Engyodontium album (fungus) |
| Molecular weight | Theoretical: 28.958791 KDa |
| Recombinant expression | Organism:  Engyodontium album (fungus) Engyodontium album (fungus) |
| Sequence | String: AAQTNAPWGL ARISSTSPGT STYYYDESAG QGSCVYVIDT GIEASHPEFE GRAQMVKTYY YSSRDGNGHG THCAGTVGSR TYGVAKKTQ LFGVKVLDDN GSGQYSTIIA GMDFVASDKN NRNCPKGVVA SLSLGGGYSS SVNSAAARLQ SSGVMVAVAA G NNNADARN ...String: AAQTNAPWGL ARISSTSPGT STYYYDESAG QGSCVYVIDT GIEASHPEFE GRAQMVKTYY YSSRDGNGHG THCAGTVGSR TYGVAKKTQ LFGVKVLDDN GSGQYSTIIA GMDFVASDKN NRNCPKGVVA SLSLGGGYSS SVNSAAARLQ SSGVMVAVAA G NNNADARN YSPASEPSVC TVGASDRYDR RSSFSNYGSV LDIFGPGTDI LSTWIGGSTR SISGTSMATP HVAGLAAYLM TL GKTTAAS ACRYIADTAN KGDLSNIPFG TVNLLAYNNY QA UniProtKB: Proteinase K |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #3: SULFATE ION
| Macromolecule | Name: SULFATE ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: SO4 |
|---|---|
| Molecular weight | Theoretical: 96.063 Da |
| Chemical component information |  ChemComp-SO4: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 23 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron crystallography |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Concentration | 20 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 12 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 273.0 K / Instrument: FEI VITROBOT MARK IV |
| Details | Proteinase K purchased from Sigma. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Temperature | Min: 77.0 K / Max: 90.0 K |
| Image recording | Film or detector model: FEI CETA (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number diffraction images: 100 / Average exposure time: 2.0 sec. / Average electron dose: 0.04 e/Å2 Details: Continuous rotation from -30 to +30 with a rotation rate of 0.2 degrees per second and a readout every 3 seconds. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: DIFFRACTION / Camera length: 1700 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | This was the new CetaD. |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.75 Å / Resolution method: DIFFRACTION PATTERN/LAYERLINES |
| Molecular replacement | Software - Name: PHENIX (ver. 2.8.2) |
| Crystallography statistics | Number intensities measured: 31013 / Number structure factors: 10807 / Fourier space coverage: 88.4 / R sym: 0.39 / R merge: 0.47 / Overall phase error: 22.41 / Overall phase residual: 22.41 / Phase error rejection criteria: 0 / High resolution: 2.75 Å / Shell - Shell ID: 1 / Shell - High resolution: 2.75 Å / Shell - Low resolution: 3.1483 Å / Shell - Number structure factors: 1831 / Shell - Phase residual: 28.11 / Shell - Fourier space coverage: 88.23 / Shell - Multiplicity: 2.9 |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT / Overall B value: 21.1 / Target criteria: R |
| Output model |  PDB-6n4u: |
 Movie
Movie Controller
Controller




 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)






















