+Search query
-Structure paper
| Title | Structural-guided identification of two modulators of β-1,3-glucan synthase FKS1. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 17, Issue 1, Page 591, Year 2025 |
| Publish date | Dec 7, 2025 |
 Authors Authors | Jialu Li / Jian Li / Angqi Zhu / Xinli Dai / Juxiu Liu / Huayi Liu / Zhili Xia / Yuefan Dong / Wenyu Qian / Lunzhi Dai / Li Guo / Chuangye Yan / Dong Deng / Yunzi Luo / Xiang Wang /  |
| PubMed Abstract | FKS1 is a β-1,3-glucan synthase critical for fungal cell wall formation and a target for antifungal drugs such as echinocandin and ibrexafungerp. However, the mechanisms regulating FKS1 activity ...FKS1 is a β-1,3-glucan synthase critical for fungal cell wall formation and a target for antifungal drugs such as echinocandin and ibrexafungerp. However, the mechanisms regulating FKS1 activity remain largely unknown. Here, we reveal that transfer RNA (tRNA) acts as an endogenous inhibitor, whereas GSR1 functions as a stabilizer of FKS1. The cryo-EM structure of FKS1 adopts a tRNA-mediated homodimer configuration, representing a quiescent state of β-1,3-glucan synthase. Unexpectedly, the copurified endogenous tRNA is identified as a potent inhibitor that suppresses FKS1 activity. Moreover, high-resolution cryo-EM density analysis enable the identification of GSR1 as an additional binding partner of FKS1. Mutagenesis experiments confirm the interaction between FKS1 and GSR1. Evolutionarily conserved GSR1 is found to increase the stability of FKS1 in β-1,3-glucan biosynthesis. Collectively, our findings identify both tRNA and GSR1 as intrinsic modulators of β-1,3-glucan biosynthesis, thereby providing opportunities for the further development of FKS1-targeted antifungal drugs. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:41354657 / PubMed:41354657 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.72 - 2.96 Å |
| Structure data | EMDB-64501, PDB-9utu: EMDB-64502, PDB-9utw: |
| Chemicals |  ChemComp-MG: 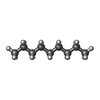 ChemComp-DD9:  ChemComp-D10: 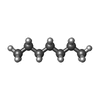 ChemComp-HP6: 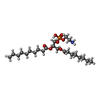 ChemComp-XKP:  ChemComp-PEF:  ChemComp-PLM:  ChemComp-DCR: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / MEMBRANE PROTEIN-RNA COMPLEX |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers








