+Search query
-Structure paper
| Title | Context-dependent translation inhibition as a cancer therapeutic modality. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 17, Issue 1, Year 2026 |
| Publish date | Feb 24, 2026 |
 Authors Authors | Paige D Diamond / Paul V Sauer / Mikael Holm / Canessa J Swanson-Swett / Lucas Ferguson / Natalie M Bratset / Grant W Wienker / Justin Seiwert Sim / Hailey K Adams / Lillian Kenner / Margot Meyers / David Gygi / Zef A Könst / Sogole Sami Bahmanyar / Lawrence G Hamann / Anthony P Schuller /  |
| PubMed Abstract | Recent work has demonstrated that some bacterial antibiotics that inhibit protein synthesis by binding the peptidyl transferase center (PTC) of the ribosome act in a context-dependent manner, ...Recent work has demonstrated that some bacterial antibiotics that inhibit protein synthesis by binding the peptidyl transferase center (PTC) of the ribosome act in a context-dependent manner, inhibiting translation elongation only at specific amino acids. However, this phenomenon has yet to be documented for compounds that inhibit the PTC of the human ribosome. Here, we use structure-based design to guide the synthesis of such PTC-binding, context-dependent inhibitors of the human ribosome, termed interdictors. In the PTC, these compounds preferentially interact with nascent protein residues that exhibit complementary physiochemical properties to the moieties of the small molecule, causing structural rearrangements in both the nascent polypeptide chain and ribosomal RNA. Further, the compounds differentially impact ribosome surveillance pathways, including the ribotoxic stress response. Finally, we confirm their anti-tumor activity after oral dosing in a mouse xenograft model of triple-negative breast cancer. Together, our data establish targeting oncogenic dependency factors through context-dependent inhibition of translation as a potential small molecule therapeutic modality for historically difficult to address cancers. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:41735331 / PubMed:41735331 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 1.9 - 2.2 Å |
| Structure data | EMDB-70083, PDB-9o3v: EMDB-70084, PDB-9o3w: EMDB-70086, PDB-9o3y: |
| Chemicals |  ChemComp-MG:  ChemComp-K:  ChemComp-SPD: 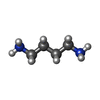 ChemComp-PUT:  ChemComp-TRS:  ChemComp-ZN:  ChemComp-HOH:  PDB-1b75:  PDB-1b76: |
| Source |
|
 Keywords Keywords | TRANSLATION / translation inhibitor / interdictor / ribosome / cryogenic-electron microscopy (cryo-EM) / ribotoxic stress response / cancer / MYC |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 homo sapiens (human)
homo sapiens (human)