+Search query
-Structure paper
| Title | Structural Basis for Polyproline-Mediated Ribosome Stalling and Rescue by the Translation Elongation Factor EF-P. |
|---|---|
| Journal, issue, pages | Mol Cell, Vol. 68, Issue 3, Page 515-527.e6, Year 2017 |
| Publish date | Nov 2, 2017 |
 Authors Authors | Paul Huter / Stefan Arenz / Lars V Bock / Michael Graf / Jan Ole Frister / Andre Heuer / Lauri Peil / Agata L Starosta / Ingo Wohlgemuth / Frank Peske / Jiří Nováček / Otto Berninghausen / Helmut Grubmüller / Tanel Tenson / Roland Beckmann / Marina V Rodnina / Andrea C Vaiana / Daniel N Wilson /    |
| PubMed Abstract | Ribosomes synthesizing proteins containing consecutive proline residues become stalled and require rescue via the action of uniquely modified translation elongation factors, EF-P in bacteria, or ...Ribosomes synthesizing proteins containing consecutive proline residues become stalled and require rescue via the action of uniquely modified translation elongation factors, EF-P in bacteria, or archaeal/eukaryotic a/eIF5A. To date, no structures exist of EF-P or eIF5A in complex with translating ribosomes stalled at polyproline stretches, and thus structural insight into how EF-P/eIF5A rescue these arrested ribosomes has been lacking. Here we present cryo-EM structures of ribosomes stalled on proline stretches, without and with modified EF-P. The structures suggest that the favored conformation of the polyproline-containing nascent chain is incompatible with the peptide exit tunnel of the ribosome and leads to destabilization of the peptidyl-tRNA. Binding of EF-P stabilizes the P-site tRNA, particularly via interactions between its modification and the CCA end, thereby enforcing an alternative conformation of the polyproline-containing nascent chain, which allows a favorable substrate geometry for peptide bond formation. |
 External links External links |  Mol Cell / Mol Cell /  PubMed:29100052 PubMed:29100052 |
| Methods | EM (single particle) |
| Resolution | 3.1 - 3.9 Å |
| Structure data | EMDB-3898, PDB-6enf: EMDB-3899, PDB-6enj:  EMDB-3900:  EMDB-3901:  EMDB-3902: |
| Chemicals | 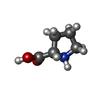 ChemComp-PRO: |
| Source |
|
 Keywords Keywords | RIBOSOME / Proline / EF-P / nascent chain / Polyproline stalled ribosome / cryo-em / elongation-factor P / stalling |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers










