+Search query
-Structure paper
| Title | Structures of protective antibodies reveal sites of vulnerability on Ebola virus. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 111, Issue 48, Page 17182-17187, Year 2014 |
| Publish date | Dec 2, 2014 |
 Authors Authors | Charles D Murin / Marnie L Fusco / Zachary A Bornholdt / Xiangguo Qiu / Gene G Olinger / Larry Zeitlin / Gary P Kobinger / Andrew B Ward / Erica Ollmann Saphire /   |
| PubMed Abstract | Ebola virus (EBOV) and related filoviruses cause severe hemorrhagic fever, with up to 90% lethality, and no treatments are approved for human use. Multiple recent outbreaks of EBOV and the likelihood ...Ebola virus (EBOV) and related filoviruses cause severe hemorrhagic fever, with up to 90% lethality, and no treatments are approved for human use. Multiple recent outbreaks of EBOV and the likelihood of future human exposure highlight the need for pre- and postexposure treatments. Monoclonal antibody (mAb) cocktails are particularly attractive candidates due to their proven postexposure efficacy in nonhuman primate models of EBOV infection. Two candidate cocktails, MB-003 and ZMAb, have been extensively evaluated in both in vitro and in vivo studies. Recently, these two therapeutics have been combined into a new cocktail named ZMapp, which showed increased efficacy and has been given compassionately to some human patients. Epitope information and mechanism of action are currently unknown for most of the component mAbs. Here we provide single-particle EM reconstructions of every mAb in the ZMapp cocktail, as well as additional antibodies from MB-003 and ZMAb. Our results illuminate key and recurring sites of vulnerability on the EBOV glycoprotein and provide a structural rationale for the efficacy of ZMapp. Interestingly, two of its components recognize overlapping epitopes and compete with each other for binding. Going forward, this work now provides a basis for strategic selection of next-generation antibody cocktails against Ebola and related viruses and a model for predicting the impact of ZMapp on potential escape mutations in ongoing or future Ebola outbreaks. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:25404321 / PubMed:25404321 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 22.0 - 24.0 Å |
| Structure data | 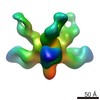 EMDB-6150: 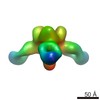 EMDB-6151: 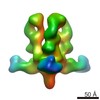 EMDB-6152: 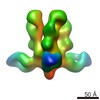 EMDB-6153: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



 Ebola virus sp.
Ebola virus sp. Homo sapiens (human)
Homo sapiens (human)