+Search query
-Structure paper
| Title | 3D cryo-electron reconstruction of BmrA, a bacterial multidrug ABC transporter in an inward-facing conformation and in a lipidic environment. |
|---|---|
| Journal, issue, pages | J Mol Biol, Vol. 426, Issue 10, Page 2059-2069, Year 2014 |
| Publish date | May 15, 2014 |
 Authors Authors | Pierre Frederic Fribourg / Mohamed Chami / Carlos Oscar S Sorzano / Francesca Gubellini / Roberto Marabini / Sergio Marco / Jean-Michel Jault / Daniel Lévy /    |
| PubMed Abstract | ABC (ATP-binding cassette) membrane exporters are efflux transporters of a wide diversity of molecule across the membrane at the expense of ATP. A key issue regarding their catalytic cycle is whether ...ABC (ATP-binding cassette) membrane exporters are efflux transporters of a wide diversity of molecule across the membrane at the expense of ATP. A key issue regarding their catalytic cycle is whether or not their nucleotide-binding domains (NBDs) are physically disengaged in the resting state. To settle this controversy, we obtained structural data on BmrA, a bacterial multidrug homodimeric ABC transporter, in a membrane-embedded state. BmrA in the apostate was reconstituted in lipid bilayers forming a mixture of ring-shaped structures of 24 or 39 homodimers. Three-dimensional models of the ring-shaped structures of 24 or 39 homodimers were calculated at 2.3 nm and 2.5 nm resolution from cryo-electron microscopy, respectively. In these structures, BmrA adopts an inward-facing open conformation similar to that found in mouse P-glycoprotein structure with the NBDs separated by 3 nm. Both lipidic leaflets delimiting the transmembrane domains of BmrA were clearly resolved. In planar membrane sheets, the NBDs were even more separated. BmrA in an ATP-bound conformation was determined from two-dimensional crystals grown in the presence of ATP and vanadate. A projection map calculated at 1.6 nm resolution shows an open outward-facing conformation. Overall, the data are consistent with a mechanism of drug transport involving large conformational changes of BmrA and show that a bacterial ABC exporter can adopt at least two open inward conformations in lipid membrane. |
 External links External links |  J Mol Biol / J Mol Biol /  PubMed:24630999 PubMed:24630999 |
| Methods | EM (single particle) |
| Resolution | 23.0 - 25.0 Å |
| Structure data |  EMDB-5525: 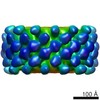 EMDB-5526: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




