+Search query
-Structure paper
| Title | Molecular mechanism of choline and ethanolamine transport in humans. |
|---|---|
| Journal, issue, pages | Nature, Vol. 630, Issue 8016, Page 501-508, Year 2024 |
| Publish date | May 22, 2024 |
 Authors Authors | Keiken Ri / Tsai-Hsuan Weng / Ainara Claveras Cabezudo / Wiebke Jösting / Yu Zhang / Andre Bazzone / Nancy C P Leong / Sonja Welsch / Raymond T Doty / Gonca Gursu / Tiffany Jia Ying Lim / Sarah Luise Schmidt / Janis L Abkowitz / Gerhard Hummer / Di Wu / Long N Nguyen / Schara Safarian /    |
| PubMed Abstract | Human feline leukaemia virus subgroup C receptor-related proteins 1 and 2 (FLVCR1 and FLVCR2) are members of the major facilitator superfamily. Their dysfunction is linked to several clinical ...Human feline leukaemia virus subgroup C receptor-related proteins 1 and 2 (FLVCR1 and FLVCR2) are members of the major facilitator superfamily. Their dysfunction is linked to several clinical disorders, including PCARP, HSAN and Fowler syndrome. Earlier studies concluded that FLVCR1 may function as a haem exporter, whereas FLVCR2 was suggested to act as a haem importer, yet conclusive biochemical and detailed molecular evidence remained elusive for the function of both transporters. Here, we show that FLVCR1 and FLVCR2 facilitate the transport of choline and ethanolamine across the plasma membrane, using a concentration-driven substrate translocation process. Through structural and computational analyses, we have identified distinct conformational states of FLVCRs and unravelled the coordination chemistry underlying their substrate interactions. Fully conserved tryptophan and tyrosine residues form the binding pocket of both transporters and confer selectivity for choline and ethanolamine through cation-π interactions. Our findings clarify the mechanisms of choline and ethanolamine transport by FLVCR1 and FLVCR2, enhance our comprehension of disease-associated mutations that interfere with these vital processes and shed light on the conformational dynamics of these major facilitator superfamily proteins during the transport cycle. |
 External links External links |  Nature / Nature /  PubMed:38778100 / PubMed:38778100 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.6 - 3.1 Å |
| Structure data | EMDB-18334, PDB-8qcs: EMDB-18335, PDB-8qct: EMDB-18336, PDB-8qcx: EMDB-18337, PDB-8qcy: EMDB-18339, PDB-8qd0: EMDB-19009, PDB-8r8t: |
| Chemicals | 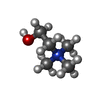 ChemComp-CHT: 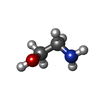 ChemComp-ETA: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / MFS / choline / human transporter / choline transport / ethanolamine transport |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers















 homo sapiens (human)
homo sapiens (human)