+検索条件
-Structure paper
| タイトル | Structure determination of inactive-state GPCRs with a universal nanobody. |
|---|---|
| ジャーナル・号・ページ | Nat Struct Mol Biol, Vol. 29, Issue 12, Page 1188-1195, Year 2022 |
| 掲載日 | 2022年11月17日 |
 著者 著者 | Michael J Robertson / Makaía M Papasergi-Scott / Feng He / Alpay B Seven / Justin G Meyerowitz / Ouliana Panova / Maria Claudia Peroto / Tao Che / Georgios Skiniotis /  |
| PubMed 要旨 | Cryogenic electron microscopy (cryo-EM) has widened the field of structure-based drug discovery by allowing for routine determination of membrane protein structures previously intractable. Despite ...Cryogenic electron microscopy (cryo-EM) has widened the field of structure-based drug discovery by allowing for routine determination of membrane protein structures previously intractable. Despite representing one of the largest classes of therapeutic targets, most inactive-state G protein-coupled receptors (GPCRs) have remained inaccessible for cryo-EM because their small size and membrane-embedded nature impedes projection alignment for high-resolution map reconstructions. Here we demonstrate that the same single-chain camelid antibody (nanobody) recognizing a grafted intracellular loop can be used to obtain cryo-EM structures of inactive-state GPCRs at resolutions comparable or better than those obtained by X-ray crystallography. Using this approach, we obtained structures of neurotensin 1 receptor bound to antagonist SR48692, μ-opioid receptor bound to alvimopan, apo somatostatin receptor 2 and histamine receptor 2 bound to famotidine. We expect this rapid, straightforward approach to facilitate the broad exploration of GPCR inactive states without the need for extensive engineering and crystallization. |
 リンク リンク |  Nat Struct Mol Biol / Nat Struct Mol Biol /  PubMed:36396979 PubMed:36396979 |
| 手法 | EM (単粒子) |
| 解像度 | 2.4 - 3.1 Å |
| 構造データ | EMDB-26589, PDB-7ul2: EMDB-26590, PDB-7ul3: EMDB-26591, PDB-7ul4: EMDB-26592, PDB-7ul5: |
| 化合物 |  ChemComp-Q6Q:  ChemComp-NA:  ChemComp-HOH:  ChemComp-FO9: 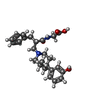 ChemComp-NG0: |
| 由来 |
|
 キーワード キーワード | MEMBRANE PROTEIN / Antagonist / Complex |
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア 万見文献について
万見文献について









 homo sapiens (ヒト)
homo sapiens (ヒト)
