+Search query
-Structure paper
| Title | Locations and in situ structure of the polymerase complex inside the virion of vesicular stomatitis virus. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 119, Issue 18, Page e2111948119, Year 2022 |
| Publish date | May 3, 2022 |
 Authors Authors | Zhu Si / Kang Zhou / Jun Tsao / Ming Luo / Z Hong Zhou /  |
| PubMed Abstract | The polymerase complex of nonsegmented negative-strand RNA viruses primarily consists of a large (L) protein and a phosphoprotein (P). L is a multifunctional enzyme carrying out RNA-dependent RNA ...The polymerase complex of nonsegmented negative-strand RNA viruses primarily consists of a large (L) protein and a phosphoprotein (P). L is a multifunctional enzyme carrying out RNA-dependent RNA polymerization and all other steps associated with transcription and replication, while P is the nonenzymatic cofactor, regulating the function and conformation of L. The structure of a purified vesicular stomatitis virus (VSV) polymerase complex containing L and associated P segments has been determined; however, the location and manner of the attachments of L and P within each virion are unknown, limiting our mechanistic understanding of VSV RNA replication and transcription and hindering engineering efforts of this widely used anticancer and vaccine vector. Here, we have used cryo-electron tomography to visualize the VSV virion, revealing the attachment of the ring-shaped L molecules to VSV nucleocapsid proteins (N) throughout the cavity of the bullet-shaped nucleocapsid. Subtomogram averaging and three-dimensional classification of regions containing N and the matrix protein (M) have yielded the in situ structure of the polymerase complex. On average, ∼55 polymerase complexes are packaged in each virion. The capping domain of L interacts with two neighboring N molecules through flexible attachments. P, which exists as a dimer, bridges separate N molecules and the connector and C-terminal domains of L. Our data provide the structural basis for recruitment of L to N by P in virus assembly and for flexible attachments between L and N, which allow a quick response of L in primary transcription upon cell entry. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:35476516 / PubMed:35476516 /  PubMed Central PubMed Central |
| Methods | EM (subtomogram averaging) |
| Resolution | 7.5 - 14.7 Å |
| Structure data |  EMDB-26465: In situ structure of Vesicular stomatitis virus (VSV) 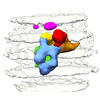 EMDB-26468: In situ structure of Vesicular stomatitis virus (VSV) polymerase complex attached to nucleoprotein (N) in VSV |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



