+Search query
-Structure paper
| Title | Complete three-dimensional structures of the Lon protease translocating a protein substrate. |
|---|---|
| Journal, issue, pages | Sci Adv, Vol. 7, Issue 42, Page eabj7835, Year 2021 |
| Publish date | Oct 15, 2021 |
 Authors Authors | Shanshan Li / Kan-Yen Hsieh / Chiao-I Kuo / Szu-Hui Lee / Grigore D Pintilie / Kaiming Zhang / Chung-I Chang /    |
| PubMed Abstract | Lon is an evolutionarily conserved proteolytic machine carrying out a wide spectrum of biological activities by degrading misfolded damaged proteins and specific cellular substrates. Lon contains a ...Lon is an evolutionarily conserved proteolytic machine carrying out a wide spectrum of biological activities by degrading misfolded damaged proteins and specific cellular substrates. Lon contains a large N-terminal domain and forms a hexameric core of fused adenosine triphosphatase and protease domains. Here, we report two complete structures of Lon engaging a substrate, determined by cryo–electron microscopy to 2.4-angstrom resolution. These structures show a multilayered architecture featuring a tensegrity triangle complex, uniquely constructed by six long N-terminal helices. The interlocked helix triangle is assembled on the top of the hexameric core to spread a web of six globular substrate-binding domains. It serves as a multipurpose platform that controls the access of substrates to the AAA+ ring, provides a ruler-based mechanism for substrate selection, and acts as a pulley device to facilitate unfolding of the translocated substrate. This work provides a complete framework for understanding the structural mechanisms of Lon. |
 External links External links |  Sci Adv / Sci Adv /  PubMed:34652947 / PubMed:34652947 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.4 Å |
| Structure data | EMDB-31534, PDB-7fd4: EMDB-31535, PDB-7fd5: |
| Chemicals |  ChemComp-AGS: 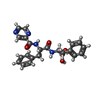 ChemComp-4KZ:  ChemComp-ADP: |
| Source |
|
 Keywords Keywords | HYDROLASE/PROTEIN BINDING / AAA+ protease / Lon / complete three-dimensional structure / N-terminal domain / CYTOSOLIC PROTEIN / HYDROLASE-PROTEIN BINDING complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers







 meiothermus taiwanensis (bacteria)
meiothermus taiwanensis (bacteria)
