+Search query
-Structure paper
| Title | Influenza hemagglutinin-specific IgA Fc-effector functionality is restricted to stalk epitopes. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 118, Issue 8, Year 2021 |
| Publish date | Feb 23, 2021 |
 Authors Authors | Alec W Freyn / Julianna Han / Jenna J Guthmiller / Mark J Bailey / Karlynn Neu / Hannah L Turner / Victoria C Rosado / Veronika Chromikova / Min Huang / Shirin Strohmeier / Sean T H Liu / Viviana Simon / Florian Krammer / Andrew B Ward / Peter Palese / Patrick C Wilson / Raffael Nachbagauer /   |
| PubMed Abstract | In this study, we utilized a panel of human immunoglobulin (Ig) IgA monoclonal antibodies isolated from the plasmablasts of eight donors after 2014/2015 influenza virus vaccination (Fluarix) to study ...In this study, we utilized a panel of human immunoglobulin (Ig) IgA monoclonal antibodies isolated from the plasmablasts of eight donors after 2014/2015 influenza virus vaccination (Fluarix) to study the binding and functional specificities of this isotype. In this cohort, isolated IgA monoclonal antibodies were primarily elicited against the hemagglutinin protein of the H1N1 component of the vaccine. To compare effector functionalities, an H1-specific subset of antibodies targeting distinct epitopes were expressed as monomeric, dimeric, or secretory IgA, as well as in an IgG1 backbone. When expressed with an IgG Fc domain, all antibodies elicited Fc-effector activity in a primary polymorphonuclear cell-based assay which differs from previous observations that found only stalk-specific antibodies activate the low-affinity FcγRIIIa. However, when expressed with IgA Fc domains, only antibodies targeting the stalk domain showed Fc-effector activity in line with these previous findings. To identify the cause of this discrepancy, we then confirmed that IgG signaling through the high-affinity FcγI receptor was not restricted to stalk epitopes. Since no corresponding high-affinity Fcα receptor exists, the IgA repertoire may therefore be limited to stalk-specific epitopes in the context of Fc receptor signaling. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:33593910 / PubMed:33593910 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.72 - 25.0 Å |
| Structure data | 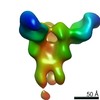 EMDB-23313:  EMDB-23314: 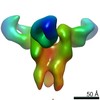 EMDB-23315: 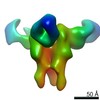 EMDB-23316:  EMDB-23317: 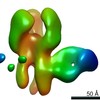 EMDB-23318:  EMDB-23319: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



 Homo sapiens (human)
Homo sapiens (human)
 Influenza A virus
Influenza A virus