+Search query
-Structure paper
| Title | Symmetrical arrangement of proteins under release-ready vesicles in presynaptic terminals. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 118, Issue 5, Year 2021 |
| Publish date | Feb 2, 2021 |
 Authors Authors | Abhijith Radhakrishnan / Xia Li / Kirill Grushin / Shyam S Krishnakumar / Jun Liu / James E Rothman /  |
| PubMed Abstract | Controlled release of neurotransmitters stored in synaptic vesicles (SVs) is a fundamental process that is central to all information processing in the brain. This relies on tight coupling of the SV ...Controlled release of neurotransmitters stored in synaptic vesicles (SVs) is a fundamental process that is central to all information processing in the brain. This relies on tight coupling of the SV fusion to action potential-evoked presynaptic Ca influx. This Ca-evoked release occurs from a readily releasable pool (RRP) of SVs docked to the plasma membrane (PM). The protein components involved in initial SV docking/tethering and the subsequent priming reactions which make the SV release ready are known. Yet, the supramolecular architecture and sequence of molecular events underlying SV release are unclear. Here, we use cryoelectron tomography analysis in cultured hippocampal neurons to delineate the arrangement of the exocytosis machinery under docked SVs. Under native conditions, we find that vesicles are initially "tethered" to the PM by a variable number of protein densities (∼10 to 20 nm long) with no discernible organization. In contrast, we observe exactly six protein masses, each likely consisting of a single SNAREpin with its bound Synaptotagmins and Complexin, arranged symmetrically connecting the "primed" vesicles to the PM. Our data indicate that the fusion machinery is likely organized into a highly cooperative framework during the priming process which enables rapid SV fusion and neurotransmitter release following Ca influx. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:33468631 / PubMed:33468631 /  PubMed Central PubMed Central |
| Methods | EM (subtomogram averaging) |
| Resolution | 44.0 Å |
| Structure data | 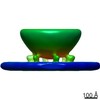 EMDB-23077: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




