+Search query
-Structure paper
| Title | Structural basis of cooling agent and lipid sensing by the cold-activated TRPM8 channel. |
|---|---|
| Journal, issue, pages | Science, Vol. 363, Issue 6430, Year 2019 |
| Publish date | Mar 1, 2019 |
 Authors Authors | Ying Yin / Son C Le / Allen L Hsu / Mario J Borgnia / Huanghe Yang / Seok-Yong Lee /  |
| PubMed Abstract | Transient receptor potential melastatin member 8 (TRPM8) is a calcium ion (Ca)-permeable cation channel that serves as the primary cold and menthol sensor in humans. Activation of TRPM8 by cooling ...Transient receptor potential melastatin member 8 (TRPM8) is a calcium ion (Ca)-permeable cation channel that serves as the primary cold and menthol sensor in humans. Activation of TRPM8 by cooling compounds relies on allosteric actions of agonist and membrane lipid phosphatidylinositol 4,5-bisphosphate (PIP), but lack of structural information has thus far precluded a mechanistic understanding of ligand and lipid sensing by TRPM8. Using cryo-electron microscopy, we determined the structures of TRPM8 in complex with the synthetic cooling compound icilin, PIP, and Ca, as well as in complex with the menthol analog WS-12 and PIP Our structures reveal the binding sites for cooling agonists and PIP in TRPM8. Notably, PIP binds to TRPM8 in two different modes, which illustrate the mechanism of allosteric coupling between PIP and agonists. This study provides a platform for understanding the molecular mechanism of TRPM8 activation by cooling agents. |
 External links External links |  Science / Science /  PubMed:30733385 / PubMed:30733385 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.4 - 4.3 Å |
| Structure data | EMDB-0487, PDB-6nr2: |
| Chemicals | 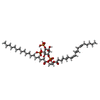 ChemComp-KXP: 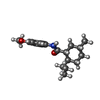 ChemComp-KXS: 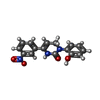 ChemComp-KX7:  ChemComp-CA: |
| Source |
|
 Keywords Keywords | TRANSPORT PROTEIN / ion channel / TRP channel / TRPM channel / TRPM8 channel / cold sensing / lipid sensing / menthol / icilin / WS-12 / PI(4 / 5)P2 / cooling agent / MEMBRANE PROTEIN / calcium-permeable ion channel / calcium-permeable channel |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 ficedula albicollis (Collared flycatcher)
ficedula albicollis (Collared flycatcher)