+Search query
-Structure paper
| Title | Functionally specific binding regions of microtubule-associated protein 2c exhibit distinct conformations and dynamics. |
|---|---|
| Journal, issue, pages | J Biol Chem, Vol. 293, Issue 34, Page 13297-13309, Year 2018 |
| Publish date | Aug 24, 2018 |
 Authors Authors | Kateřina Melková / Vojtěch Zapletal / Séverine Jansen / Erik Nomilner / Milan Zachrdla / Jozef Hritz / Jiří Nováček / Markus Zweckstetter / Malene R Jensen / Martin Blackledge / Lukáš Žídek /    |
| PubMed Abstract | Microtubule-associated protein 2c (MAP2c) is a 49-kDa intrinsically disordered protein regulating the dynamics of microtubules in developing neurons. MAP2c differs from its sequence homologue Tau in ...Microtubule-associated protein 2c (MAP2c) is a 49-kDa intrinsically disordered protein regulating the dynamics of microtubules in developing neurons. MAP2c differs from its sequence homologue Tau in the pattern and kinetics of phosphorylation by cAMP-dependent protein kinase (PKA). Moreover, the mechanisms through which MAP2c interacts with its binding partners and the conformational changes and dynamics associated with these interactions remain unclear. Here, we used NMR relaxation and paramagnetic relaxation enhancement techniques to determine the dynamics and long-range interactions within MAP2c. The relaxation rates revealed large differences in flexibility of individual regions of MAP2c, with the lowest flexibility observed in the known and proposed binding sites. Quantitative conformational analyses of chemical shifts, small-angle X-ray scattering (SAXS), and paramagnetic relaxation enhancement measurements disclosed that MAP2c regions interacting with important protein partners, including Fyn tyrosine kinase, plectin, and PKA, adopt specific conformations. High populations of polyproline II and α-helices were found in Fyn- and plectin-binding sites of MAP2c, respectively. The region binding the regulatory subunit of PKA consists of two helical motifs bridged by a more extended conformation. Of note, although MAP2c and Tau did not differ substantially in their conformations in regions of high sequence identity, we found that they differ significantly in long-range interactions, dynamics, and local conformation motifs in their N-terminal domains. These results highlight that the N-terminal regions of MAP2c provide important specificity to its regulatory roles and indicate a close relationship between MAP2c's biological functions and conformational behavior. |
 External links External links |  J Biol Chem / J Biol Chem /  PubMed:29925592 / PubMed:29925592 /  PubMed Central PubMed Central |
| Methods | SAS (X-ray synchrotron) |
| Structure data | 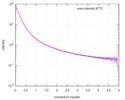 SASDD62: Microtubule associated protein MAP2c (isoform 3); 12 mg/ml 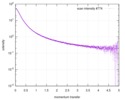 SASDD72: Microtubule associated protein MAP2c (isoform 3); 6 mg/ml 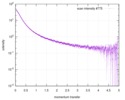 SASDD82: Microtubule associated protein MAP2c (isoform 3); 3 mg/ml 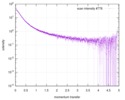 SASDD92: Microtubule associated protein MAP2c (isoform 3); 1.5 mg/ml 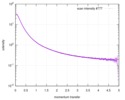 SASDDA2: Phosphorylated Microtubule associated protein MAP2c (isoform 3); 13.6 mg/ml 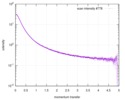 SASDDB2: Phosphorylated Microtubule associated protein MAP2c (isoform 3); 6.8 mg/ml 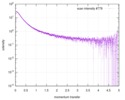 SASDDC2: Phosphorylated Microtubule associated protein MAP2c (isoform 3); 1.7 mg/ml |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




