-Search query
-Search result
Showing 1 - 50 of 336 items for (author: chou & t)

EMDB-39546: 
SARS-CoV-2 Delta Spike in complex with JL-8C
Method: single particle / : Nguyen VHT, Chen X

EMDB-39547: 
SARS-CoV-2 Delta Spike in complex with JM-1A
Method: single particle / : Nguyen VHT, Chen X

EMDB-39685: 
SARS-CoV-2 Delta Spike in complex with Fab of JE-5C
Method: single particle / : Chen X, Wu YM

EMDB-39686: 
SARS-CoV-2 Spike (BA.1) in complex with Fab of JH-8B
Method: single particle / : Chen X, Wu YM
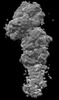
EMDB-44479: 
Cryo-EM structure of synthetic claudin-4 complex with Clostridium perfringens enterotoxin C-terminal domain, sFab COP-2, and Nanobody
Method: single particle / : Vecchio AJ

EMDB-36800: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-36801: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state, focused refined on KtrA octamer
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-36802: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state, focused refined on KtrB dimer
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-36803: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of MgCl2
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-36804: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-38477: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, vertical C2 symmetry axis
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-38478: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, C1 symmetry
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

PDB-8k1s: 
Potassium transporter KtrAB from Bacillus subtilis in ADP-bound state
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

PDB-8k1t: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of MgCl2
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

PDB-8k1u: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

PDB-8xmh: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, vertical C2 symmetry axis
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

PDB-8xmi: 
Potassium transporter KtrAB from Bacillus subtilis in ATP-bound state with addition of EDTA and EGTA, C1 symmetry
Method: single particle / : Chang YK, Chiang WT, Hu NJ, Tsai MD

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vye: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyf: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyg: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-41423: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41424: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41425: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41777: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41778: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41779: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

PDB-8tnp: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

PDB-8tnq: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

PDB-8tnr: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-19431: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free)
Method: single particle / : Mazza T, Beis K

EMDB-19433: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid
Method: single particle / : Mazza T, Beis K

PDB-8rq3: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free)
Method: single particle / : Mazza T, Beis K

PDB-8rq4: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid
Method: single particle / : Mazza T, Beis K

EMDB-40811: 
Cryo-EM structure of NLRP3 open octamer
Method: single particle / : Yu X, Matico RE, Miller R, Schoubroeck BV, Grauwen K, Suarez J, Pietrak B, Haloi N, Yin Y, Tresadern GJ, Perez-Benito L, Lindahl E, Bottelbergs A, Oehlrich D, Opdenbosch NV, Sharma S
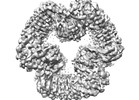
EMDB-40820: 
Cryo-EM structure of NLRP3 closed hexamer
Method: single particle / : Yu X, Matico RE, Miller R, Schoubroeck BV, Grauwen K, Suarez J, Pietrak B, Haloi N, Yin Y, Tresadern GJ, Perez-Benito L, Lindahl E, Bottelbergs A, Oehlrich D, Opdenbosch NV, Sharma S
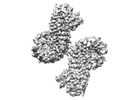
EMDB-40855: 
Structure of NLRP3 and NEK7 complex
Method: single particle / : Yu X, Matico RE, Miller R, Schoubroeck BV, Grauwen K, Suarez J, Pietrak B, Haloi N, Yin Y, Tresadern GJ, Perez-Benito L, Lindahl E, Bottelbergs A, Oehlrich D, Opdenbosch NV, Sharma S

PDB-8swf: 
Cryo-EM structure of NLRP3 open octamer
Method: single particle / : Yu X, Matico RE, Miller R, Schoubroeck BV, Grauwen K, Suarez J, Pietrak B, Haloi N, Yin Y, Tresadern GJ, Perez-Benito L, Lindahl E, Bottelbergs A, Oehlrich D, Opdenbosch NV, Sharma S

PDB-8swk: 
Cryo-EM structure of NLRP3 closed hexamer
Method: single particle / : Yu X, Matico RE, Miller R, Schoubroeck BV, Grauwen K, Suarez J, Pietrak B, Haloi N, Yin Y, Tresadern GJ, Perez-Benito L, Lindahl E, Bottelbergs A, Oehlrich D, Opdenbosch NV, Sharma S

PDB-8sxn: 
Structure of NLRP3 and NEK7 complex
Method: single particle / : Yu X, Matico RE, Miller R, Schoubroeck BV, Grauwen K, Suarez J, Pietrak B, Haloi N, Yin Y, Tresadern GJ, Perez-Benito L, Lindahl E, Bottelbergs A, Oehlrich D, Opdenbosch NV, Sharma S

EMDB-42787: 
Arp2/3 branch junction complex, ADP state
Method: single particle / : Chavali SS, Chou SZ, Sindelar CV

EMDB-42788: 
Arp2/3 branch junction complex, BeFx state
Method: single particle / : Chavali SS, Chou SZ, Sindelar CV

EMDB-42829: 
Straight actin filament from Arp2/3 branch junction sample (ADP)
Method: helical / : Chavali SS, Chou SZ, Sindelar CV

EMDB-42830: 
Straight actin filament from Arp2/3 branch junction sample (ADP-BeFx)
Method: helical / : Chavali SS, Chou SZ, Sindelar CV

PDB-8uxw: 
Arp2/3 branch junction complex, ADP state
Method: single particle / : Chavali SS, Chou SZ, Sindelar CV

PDB-8uxx: 
Arp2/3 branch junction complex, BeFx state
Method: single particle / : Chavali SS, Chou SZ, Sindelar CV

PDB-8uz0: 
Straight actin filament from Arp2/3 branch junction sample (ADP)
Method: helical / : Chavali SS, Chou SZ, Sindelar CV

PDB-8uz1: 
Straight actin filament from Arp2/3 branch junction sample (ADP-BeFx)
Method: helical / : Chavali SS, Chou SZ, Sindelar CV
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
