[English] 日本語
 Yorodumi
Yorodumi- EMDB-7015: Structure of 30S ribosomal subunit and RNA polymerase complex in ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7015 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
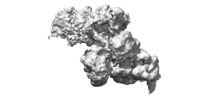
| Title | Structure of 30S ribosomal subunit and RNA polymerase complex in rotated state | ||||||||||||
 Map data Map data | Structure of 30S ribosomal subunit and RNA polymerase complex in rotated state | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | 30S subunit RNA polymerase complex / RIBOSOME | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationDNA-directed RNA polymerase complex / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / tRNA binding ...DNA-directed RNA polymerase complex / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / tRNA binding / protein dimerization activity / rRNA binding / ribosome / structural constituent of ribosome / translation / ribonucleoprotein complex / DNA-templated transcription / mRNA binding / magnesium ion binding / DNA binding / RNA binding / zinc ion binding / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.9 Å | ||||||||||||
 Authors Authors | Demo G / Rasouly A | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: Structure of RNA polymerase bound to ribosomal 30S subunit. Authors: Gabriel Demo / Aviram Rasouly / Nikita Vasilyev / Vladimir Svetlov / Anna B Loveland / Ruben Diaz-Avalos / Nikolaus Grigorieff / Evgeny Nudler / Andrei A Korostelev /  Abstract: In bacteria, mRNA transcription and translation are coupled to coordinate optimal gene expression and maintain genome stability. Coupling is thought to involve direct interactions between RNA ...In bacteria, mRNA transcription and translation are coupled to coordinate optimal gene expression and maintain genome stability. Coupling is thought to involve direct interactions between RNA polymerase (RNAP) and the translational machinery. We present cryo-EM structures of RNAP core bound to the small ribosomal 30S subunit. The complex is stable under cell-like ionic conditions, consistent with functional interaction between RNAP and the 30S subunit. The RNA exit tunnel of RNAP aligns with the Shine-Dalgarno-binding site of the 30S subunit. Ribosomal protein S1 forms a wall of the tunnel between RNAP and the 30S subunit, consistent with its role in directing mRNAs onto the ribosome. The nucleic-acid-binding cleft of RNAP samples distinct conformations, suggesting different functional states during transcription-translation coupling. The architecture of the 30S•RNAP complex provides a structural basis for co-localization of the transcriptional and translational machineries, and inform future mechanistic studies of coupled transcription and translation. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7015.map.gz emd_7015.map.gz | 29.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7015-v30.xml emd-7015-v30.xml emd-7015.xml emd-7015.xml | 48 KB 48 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7015.png emd_7015.png | 87.2 KB | ||
| Filedesc metadata |  emd-7015.cif.gz emd-7015.cif.gz | 11.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7015 http://ftp.pdbj.org/pub/emdb/structures/EMD-7015 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7015 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7015 | HTTPS FTP |
-Validation report
| Summary document |  emd_7015_validation.pdf.gz emd_7015_validation.pdf.gz | 496.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_7015_full_validation.pdf.gz emd_7015_full_validation.pdf.gz | 495.7 KB | Display | |
| Data in XML |  emd_7015_validation.xml.gz emd_7015_validation.xml.gz | 6.7 KB | Display | |
| Data in CIF |  emd_7015_validation.cif.gz emd_7015_validation.cif.gz | 7.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7015 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7015 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7015 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7015 | HTTPS FTP |
-Related structure data
| Related structure data |  6awcMC  7014C  7016C  6awbC  6awdC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7015.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7015.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of 30S ribosomal subunit and RNA polymerase complex in rotated state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.6428 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : 30S ribosomal subunit and RNA polymerase complex
+Supramolecule #1: 30S ribosomal subunit and RNA polymerase complex
+Supramolecule #2: 30S ribosomal subunit
+Supramolecule #3: RNA polymerase
+Macromolecule #1: 16S rRNA
+Macromolecule #2: DNA-directed RNA polymerase subunit alpha
+Macromolecule #3: DNA-directed RNA polymerase subunit beta
+Macromolecule #4: DNA-directed RNA polymerase subunit beta'
+Macromolecule #5: DNA-directed RNA polymerase subunit omega
+Macromolecule #6: 30S ribosomal protein S1
+Macromolecule #7: 30S ribosomal protein S2
+Macromolecule #8: 30S ribosomal protein S3
+Macromolecule #9: 30S ribosomal protein S4
+Macromolecule #10: 30S ribosomal protein S5
+Macromolecule #11: 30S ribosomal protein S6
+Macromolecule #12: 30S ribosomal protein S7
+Macromolecule #13: 30S ribosomal protein S8
+Macromolecule #14: 30S ribosomal protein S9
+Macromolecule #15: 30S ribosomal protein S10
+Macromolecule #16: 30S ribosomal protein S11
+Macromolecule #17: 30S ribosomal protein S12
+Macromolecule #18: 30S ribosomal protein S13
+Macromolecule #19: 30S ribosomal protein S14
+Macromolecule #20: 30S ribosomal protein S15
+Macromolecule #21: 30S ribosomal protein S16
+Macromolecule #22: 30S ribosomal protein S17
+Macromolecule #23: 30S ribosomal protein S18
+Macromolecule #24: 30S ribosomal protein S19
+Macromolecule #25: 30S ribosomal protein S20
+Macromolecule #26: 30S ribosomal protein S21
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 Details: 20 mM Tris-HCl, pH 7.0, 100 mM NH4Cl, 10 mM MgCl2, 0.5 mM EDTA, 6 mM BME |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV Details: 2.5 uL of 50 nM 30S and 150 nM RNAP was applied to the grid. After a 30 second incubation, the grid was blotted for 5 seconds at blotting power 8.. |
| Details | 50 nM 30S, 150 nM RNAP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-50 / Number grids imaged: 1 / Number real images: 2527 / Average electron dose: 40.0 e/Å2 Details: The average electron dose 40.0 (e-/A2) represents the total dose for one collected movie. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 25000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 500 / Target criteria: Correlation Coefficient |
|---|---|
| Output model |  PDB-6awc: |
 Movie
Movie Controller
Controller