[English] 日本語
 Yorodumi
Yorodumi- EMDB-1288: Asymmetric binding of transferrin receptor to parvovirus capsids. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1288 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Asymmetric binding of transferrin receptor to parvovirus capsids. | |||||||||
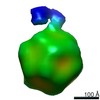
 Map data Map data | Three-dimensional reconstruction of canine parvovirus with bound feline transferrin receptor. | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationtransferrin receptor activity / negative regulation of mitochondrial fusion / transferrin transport / Transferrin endocytosis and recycling / positive regulation of isotype switching / Differentiation of keratinocytes in interfollicular epidermis in mammalian skin / response to copper ion / response to iron ion / RND1 GTPase cycle / response to manganese ion ...transferrin receptor activity / negative regulation of mitochondrial fusion / transferrin transport / Transferrin endocytosis and recycling / positive regulation of isotype switching / Differentiation of keratinocytes in interfollicular epidermis in mammalian skin / response to copper ion / response to iron ion / RND1 GTPase cycle / response to manganese ion / RND2 GTPase cycle / positive regulation of bone resorption / RHOB GTPase cycle / Golgi Associated Vesicle Biogenesis / RHOC GTPase cycle / RHOJ GTPase cycle / RHOQ GTPase cycle / CDC42 GTPase cycle / RHOH GTPase cycle / RHOG GTPase cycle / transport across blood-brain barrier / RHOA GTPase cycle / RAC2 GTPase cycle / RAC3 GTPase cycle / response to nutrient / response to retinoic acid / positive regulation of T cell proliferation / clathrin-coated pit / positive regulation of B cell proliferation / Hsp70 protein binding / RAC1 GTPase cycle / osteoclast differentiation / clathrin-coated endocytic vesicle membrane / cellular response to leukemia inhibitory factor / acute-phase response / positive regulation of protein-containing complex assembly / HFE-transferrin receptor complex / recycling endosome / receptor internalization / positive regulation of protein localization to nucleus / cellular response to xenobiotic stimulus / recycling endosome membrane / Cargo recognition for clathrin-mediated endocytosis / positive regulation of peptidyl-serine phosphorylation / double-stranded RNA binding / extracellular vesicle / Clathrin-mediated endocytosis / virus receptor activity / melanosome / positive regulation of NF-kappaB transcription factor activity / positive regulation of canonical NF-kappaB signal transduction / iron ion transport / basolateral plasma membrane / cytoplasmic vesicle / intracellular iron ion homeostasis / blood microparticle / endosome / endosome membrane / early endosome / response to hypoxia / intracellular signal transduction / positive regulation of protein phosphorylation / external side of plasma membrane / intracellular membrane-bounded organelle / positive regulation of gene expression / negative regulation of apoptotic process / protein-containing complex binding / protein kinase binding / perinuclear region of cytoplasm / cell surface / protein homodimerization activity / RNA binding / extracellular space / extracellular exosome / extracellular region / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Canine parvovirus 2 Canine parvovirus 2 | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 27.0 Å | |||||||||
 Authors Authors | Hafenstein S / Palermo LM / Xiao C / Kostyuchenko VA / Morais M / Nelson CDS / Chipman PR / Bowman VD / Battisti AJ / Parrish CR / Rossmann MG | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2007 Journal: Proc Natl Acad Sci U S A / Year: 2007Title: Asymmetric binding of transferrin receptor to parvovirus capsids. Authors: Susan Hafenstein / Laura M Palermo / Victor A Kostyuchenko / Chuan Xiao / Marc C Morais / Christian D S Nelson / Valorie D Bowman / Anthony J Battisti / Paul R Chipman / Colin R Parrish / Michael G Rossmann /  Abstract: Although many viruses are icosahedral when they initially bind to one or more receptor molecules on the cell surface, such an interaction is asymmetric, probably causing a breakdown in the symmetry ...Although many viruses are icosahedral when they initially bind to one or more receptor molecules on the cell surface, such an interaction is asymmetric, probably causing a breakdown in the symmetry and conformation of the original infecting virion in preparation for membrane penetration and release of the viral genome. Cryoelectron microscopy and biochemical analyses show that transferrin receptor, the cellular receptor for canine parvovirus, can bind to only one or a few of the 60 icosahedrally equivalent sites on the virion, indicating that either canine parvovirus has inherent asymmetry or binding of receptor induces asymmetry. The asymmetry of receptor binding to canine parvovirus is reminiscent of the special portal in tailed bacteriophages and some large, icosahedral viruses. Asymmetric interactions of icosahedral viruses with their hosts might be a more common phenomenon than previously thought and may have been obscured by averaging in previous crystallographic and electron microscopic structure determinations. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1288.map.gz emd_1288.map.gz | 48 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1288-v30.xml emd-1288-v30.xml emd-1288.xml emd-1288.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
| Images |  1288.gif 1288.gif | 15.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1288 http://ftp.pdbj.org/pub/emdb/structures/EMD-1288 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1288 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1288 | HTTPS FTP |
-Validation report
| Summary document |  emd_1288_validation.pdf.gz emd_1288_validation.pdf.gz | 297.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1288_full_validation.pdf.gz emd_1288_full_validation.pdf.gz | 297.4 KB | Display | |
| Data in XML |  emd_1288_validation.xml.gz emd_1288_validation.xml.gz | 6.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1288 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1288 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1288 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1288 | HTTPS FTP |
-Related structure data
| Related structure data |  2nsuMC  1287C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1288.map.gz / Format: CCP4 / Size: 62.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1288.map.gz / Format: CCP4 / Size: 62.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Three-dimensional reconstruction of canine parvovirus with bound feline transferrin receptor. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Canine parvovirus complexed with feline transferrin receptor
| Entire | Name: Canine parvovirus complexed with feline transferrin receptor |
|---|---|
| Components |
|
-Supramolecule #1000: Canine parvovirus complexed with feline transferrin receptor
| Supramolecule | Name: Canine parvovirus complexed with feline transferrin receptor type: sample / ID: 1000 Oligomeric state: one homodimer of fTfR binds to one icosahedral CPV particle Number unique components: 2 |
|---|
-Supramolecule #1: Canine parvovirus 2
| Supramolecule | Name: Canine parvovirus 2 / type: virus / ID: 1 / Name.synonym: CPV / Details: empty capsids / NCBI-ID: 246878 / Sci species name: Canine parvovirus 2 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes / Syn species name: CPV |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 280 Å / T number (triangulation number): 1 |
-Macromolecule #1: feline transferrin receptor
| Macromolecule | Name: feline transferrin receptor / type: protein_or_peptide / ID: 1 / Name.synonym: feTfR / Details: ectodomain only / Number of copies: 2 / Oligomeric state: dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  unidentified baculovirus / Recombinant plasmid: pcDNA 3.1 unidentified baculovirus / Recombinant plasmid: pcDNA 3.1 |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 20 mM Tris-HCl |
|---|---|
| Staining | Type: NEGATIVE / Details: no staining |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Temperature | Average: 100 K |
| Date | Jan 22, 2003 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 14 µm / Number real images: 115 / Average electron dose: 25.96 e/Å2 / Bits/pixel: 8 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 54000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 3.9 µm / Nominal defocus min: 1.7 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: 626 Single Tilt Cryotransfer System / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: CTF correction of each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 27.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: XMIPP with modifications Details: Intermediate reconstructions were made from images with virus density subtracted, final reconstruction was calculated from complete images. Number images used: 8566 |
| Final angle assignment | Details: initial angles were selected using SPIDER VO EA procedure with limits for theta of 0-40, phi of 0-72 to cover an asymmetric unit of an icosahedron; During orientation search the angles were ...Details: initial angles were selected using SPIDER VO EA procedure with limits for theta of 0-40, phi of 0-72 to cover an asymmetric unit of an icosahedron; During orientation search the angles were changed to be one of the 60 icosahedral symmetry equivalent triplet. |
| Final two d classification | Number classes: 226 |
 Movie
Movie Controller
Controller