+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3582 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
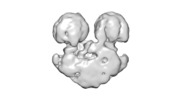
| Title | ATP synthase dimer from Tetrahymena thermophila | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |   Tetrahymena (eukaryote) Tetrahymena (eukaryote) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  negative staining / Resolution: 20.0 Å negative staining / Resolution: 20.0 Å | |||||||||
 Authors Authors | Dudkina NV / Chaban Y | |||||||||
 Citation Citation |  Journal: Biochim Biophys Acta / Year: 2014 Journal: Biochim Biophys Acta / Year: 2014Title: Structures of mitochondrial oxidative phosphorylation supercomplexes and mechanisms for their stabilisation. Authors: Yuriy Chaban / Egbert J Boekema / Natalya V Dudkina /   Abstract: Oxidative phosphorylation (OXPHOS) is the main source of energy in eukaryotic cells. This process is performed by means of electron flow between four enzymes, of which three are proton pumps, in the ...Oxidative phosphorylation (OXPHOS) is the main source of energy in eukaryotic cells. This process is performed by means of electron flow between four enzymes, of which three are proton pumps, in the inner mitochondrial membrane. The energy accumulated in the proton gradient over the inner membrane is utilized for ATP synthesis by a fifth OXPHOS complex, ATP synthase. Four of the OXPHOS protein complexes associate into stable entities called respiratory supercomplexes. This review summarises the current view on the arrangement of the electron transport chain in mitochondrial cristae. The functional role of the supramolecular organisation of the OXPHOS system and the factors that stabilise such organisation are highlighted. This article is part of a Special Issue entitled: Dynamic and ultrastructure of bioenergetic membranes and their components. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3582.map.gz emd_3582.map.gz | 805.5 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3582-v30.xml emd-3582-v30.xml emd-3582.xml emd-3582.xml | 8.2 KB 8.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3582.png emd_3582.png | 51.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3582 http://ftp.pdbj.org/pub/emdb/structures/EMD-3582 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3582 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3582 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3582.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3582.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 3.8 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ATP synthase dimer
| Entire | Name: ATP synthase dimer |
|---|---|
| Components |
|
-Supramolecule #1: ATP synthase dimer
| Supramolecule | Name: ATP synthase dimer / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Tetrahymena (eukaryote) Tetrahymena (eukaryote) |
| Molecular weight | Theoretical: 600 kDa/nm |
-Experimental details
-Structure determination
| Method |  negative staining negative staining |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Staining | Type: NEGATIVE / Material: Uranyl acetate / Details: 2% water solution |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM12 |
|---|---|
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 40.0 e/Å2 |
- Image processing
Image processing
| Particle selection | Number selected: 50000 |
|---|---|
| CTF correction | Software: (Name: IMAGIC, EMAN, SPIDER) |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: IMAGIC |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: IMAGIC |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: FSC 0.5 CUT-OFF / Number images used: 6000 |
 Movie
Movie Controller
Controller