+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of apo-GPR55-G13 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / G protein / cryo-EM / membrane protein / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcannabinoid receptor activity / Class A/1 (Rhodopsin-like receptors) / negative regulation of osteoclast differentiation / positive regulation of Rho protein signal transduction / bone resorption / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta ...cannabinoid receptor activity / Class A/1 (Rhodopsin-like receptors) / negative regulation of osteoclast differentiation / positive regulation of Rho protein signal transduction / bone resorption / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / fibroblast proliferation / Ca2+ pathway / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Ras protein signal transduction / positive regulation of ERK1 and ERK2 cascade / cell population proliferation / Extra-nuclear estrogen signaling / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / synapse / protein-containing complex binding / signal transduction / extracellular exosome / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.01 Å | |||||||||
 Authors Authors | Hua T / Liu ZJ / Cherezov V / Chang H / Li XT / Shen L | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2025 Journal: Cell Res / Year: 2025Title: Structure basis of ligand recognition and activation of GPR55. Authors: Hao Chang / Xiaoting Li / Ling Shen / Xuanrui Ge / Shuming Hao / Lijie Wu / Shenhui Liu / Junlin Liu / Vadim Cherezov / Tian Hua /   | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60990.map.gz emd_60990.map.gz | 57 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60990-v30.xml emd-60990-v30.xml emd-60990.xml emd-60990.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
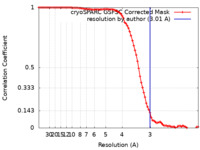
| FSC (resolution estimation) |  emd_60990_fsc.xml emd_60990_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_60990.png emd_60990.png | 57.2 KB | ||
| Filedesc metadata |  emd-60990.cif.gz emd-60990.cif.gz | 6.5 KB | ||
| Others |  emd_60990_half_map_1.map.gz emd_60990_half_map_1.map.gz emd_60990_half_map_2.map.gz emd_60990_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60990 http://ftp.pdbj.org/pub/emdb/structures/EMD-60990 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60990 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60990 | HTTPS FTP |
-Related structure data
| Related structure data |  9iy8MC  9iyaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60990.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60990.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_60990_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_60990_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of GPR55 with Guanine nucleotide-binding protein ...
| Entire | Name: Ternary complex of GPR55 with Guanine nucleotide-binding protein and nanobody |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of GPR55 with Guanine nucleotide-binding protein ...
| Supramolecule | Name: Ternary complex of GPR55 with Guanine nucleotide-binding protein and nanobody type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 175 KDa |
-Macromolecule #1: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.045629 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: IGRARGFSEL DQLRQEAEQL KNQIRDARKA CADATLSQIT NNIDPVGRIQ MRTRRTLRGH LAKIYAMHWG TDSRLLVSAS QDGKLIIWD SYTTNKVHAI PLRSSWVMTC AYAPSGNYVA CGGLDNICSI YNLKTREGNV RVSRELAGHT GYLSCCRFLD D NQIVTSSG ...String: IGRARGFSEL DQLRQEAEQL KNQIRDARKA CADATLSQIT NNIDPVGRIQ MRTRRTLRGH LAKIYAMHWG TDSRLLVSAS QDGKLIIWD SYTTNKVHAI PLRSSWVMTC AYAPSGNYVA CGGLDNICSI YNLKTREGNV RVSRELAGHT GYLSCCRFLD D NQIVTSSG DTTCALWDIE TGQQTTTFTG HTGDVMSLSL APDTRLFVSG ACDASAKLWD VREGMCRQTF TGHESDINAI CF FPNGNAF ATGSDDATCR LFDLRADQEL MTYSHDNIIC GITSVSFSKS GRLLLAGYDD FNCNVWDALK ADRAGVLAGH DNR VSCLGV TDDGMAVATG SWDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.729947 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ASNNTASIAQ ARKLVEQLKM EANIDRIKVS KAAADLMAYC EAHAKEDPLL TPVPASENPF REKKFFCAIL UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #3: G-protein coupled receptor 55
| Macromolecule | Name: G-protein coupled receptor 55 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 36.584254 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SQQNTSGDCL FDGVNELMKT LQFAVHIPTF VLGLLLNLLA IHGFSTFLKN RWPDYAATSI YMINLAVFDL LLVLSLPFKM VLSQVQSPF PSLCTLVECL YFVSMYGSVF TRCFISMDRF LAIRYPLLVS HLRSPRKIFG ICCTIWVLVW TGSIPIYSFH G KVEKYMCF ...String: SQQNTSGDCL FDGVNELMKT LQFAVHIPTF VLGLLLNLLA IHGFSTFLKN RWPDYAATSI YMINLAVFDL LLVLSLPFKM VLSQVQSPF PSLCTLVECL YFVSMYGSVF TRCFISMDRF LAIRYPLLVS HLRSPRKIFG ICCTIWVLVW TGSIPIYSFH G KVEKYMCF HNMSDDTWSA KVFFPLEVFG FLLPMGIMGF CCSRSIHILL GRRDHTQDWV QQKACIYSIA ASLAVFVVSF LP VHLGFFL QFLVRNSFIV ECRAKQSISF FLQLSMCFSN VNCCLDVFCY YFVIKEFRMN IRAHRPSRVQ LVLQDTTISR G UniProtKB: G-protein coupled receptor 55 |
-Macromolecule #4: scFV16
| Macromolecule | Name: scFV16 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.709748 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGSSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGSSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAA |
-Macromolecule #5: Guanine nucleotide-binding protein subunit alpha-13
| Macromolecule | Name: Guanine nucleotide-binding protein subunit alpha-13 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.588428 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTLSAEDK AAAERSKEID KCLSREKTYV KRLVKILLLG ADNSGKSTFL KQMRIIHGGS GGSGGTKGIH EYDFEIKNVP FKMVDVGGQ RSERKRWFEC FDSVTSILFL VDSSDFNRLT ESLNDFETIV NNRVFSNVSI ILFLNKTDLL EEKVQIVSIK D YFLEFEGD ...String: MGSTLSAEDK AAAERSKEID KCLSREKTYV KRLVKILLLG ADNSGKSTFL KQMRIIHGGS GGSGGTKGIH EYDFEIKNVP FKMVDVGGQ RSERKRWFEC FDSVTSILFL VDSSDFNRLT ESLNDFETIV NNRVFSNVSI ILFLNKTDLL EEKVQIVSIK D YFLEFEGD PHCLRDVQKF LVECFRNKRR DQQQKPLYHH FTTAINTENA RLIFRDVKDT ILHDNLKQLM LQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT / Overall B value: 107.75 |
|---|---|
| Output model |  PDB-9iy8: |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)