[English] 日本語
 Yorodumi
Yorodumi- EMDB-5783: Bacteriophage-encoded Tubulin Homologue PhuZ Forms a Three-Strand... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5783 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
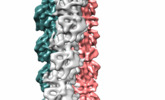
| Title | Bacteriophage-encoded Tubulin Homologue PhuZ Forms a Three-Stranded Filament | |||||||||
 Map data Map data | Reconstruction of PhuZ filament structure | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PhuZ / FtsZ/tubulin-related protein / filament / helical reconstruction / Pseudomonas bacteriophages / 201phi2-1 | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of viral life cycle / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / host cell cytoplasm / GTPase activity / GTP binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Pseudomonas phage 201phi2-1 (virus) Pseudomonas phage 201phi2-1 (virus) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 7.1 Å | |||||||||
 Authors Authors | Zehr EA / Kraemer JA / Erb ML / Coker JKC / Montabana EA / Pogliano J / Agard DA | |||||||||
 Citation Citation |  Journal: Structure / Year: 2014 Journal: Structure / Year: 2014Title: The structure and assembly mechanism of a novel three-stranded tubulin filament that centers phage DNA. Authors: Elena A Zehr / James A Kraemer / Marcella L Erb / Joanna K C Coker / Elizabeth A Montabana / Joe Pogliano / David A Agard /  Abstract: Tubulins are a universally conserved protein superfamily that carry out diverse biological roles by assembling filaments with very different architectures. The underlying basis of this structural ...Tubulins are a universally conserved protein superfamily that carry out diverse biological roles by assembling filaments with very different architectures. The underlying basis of this structural diversity is poorly understood. Here, we determine a 7.1 Å cryo-electron microscopy reconstruction of the bacteriophage-encoded PhuZ filament and provide molecular-level insight into its cooperative assembly mechanism. The PhuZ family of tubulins is required to actively center the phage within infected host cells, facilitating efficient phage replication. Our reconstruction and derived model reveal the first example of a three-stranded tubulin filament. We show that the elongated C-terminal tail simultaneously stabilizes both longitudinal and lateral interactions, which in turn define filament architecture. Identified interaction surfaces are conserved within the PhuZ family, and their mutagenesis compromises polymerization in vitro and in vivo. Combining kinetic modeling of PhuZ filament assembly and structural data, we suggest a common filament structure and assembly mechanism for the PhuZ family of tubulins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5783.map.gz emd_5783.map.gz | 28.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5783-v30.xml emd-5783-v30.xml emd-5783.xml emd-5783.xml | 10.7 KB 10.7 KB | Display Display |  EMDB header EMDB header |
| Images |  400_5783.gif 400_5783.gif 80_5783.gif 80_5783.gif | 75.5 KB 5.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5783 http://ftp.pdbj.org/pub/emdb/structures/EMD-5783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5783 | HTTPS FTP |
-Related structure data
| Related structure data |  3j5vMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5783.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5783.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of PhuZ filament structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.20398 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PhuZ201 filament
| Entire | Name: PhuZ201 filament |
|---|---|
| Components |
|
-Supramolecule #1000: PhuZ201 filament
| Supramolecule | Name: PhuZ201 filament / type: sample / ID: 1000 / Oligomeric state: filament / Number unique components: 1 |
|---|
-Macromolecule #1: PhuZ
| Macromolecule | Name: PhuZ / type: protein_or_peptide / ID: 1 / Name.synonym: PhuZ201 / Oligomeric state: polymer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage 201phi2-1 (virus) / synonym: bacteriophage Pseudomonas phage 201phi2-1 (virus) / synonym: bacteriophage |
| Molecular weight | Experimental: 34.621 KDa / Theoretical: 34.621 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Phage tubulin-like protein / InterPro: Tubulin/FtsZ, GTPase domain |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 50 mM HEPES, 125 mM KCl, 5 mM MgCl2, 5% glycerol, 1 mM GMPCPP |
| Grid | Details: C-FLAT holey carbon grid, 200 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: FEI VITROBOT MARK III Method: Sample was applied to C-FLAT holey carbon grids, blotted for 2 seconds, and flash frozen in liquid ethane |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Astigmatism corrected at 250,000 times magnification |
| Date | Nov 21, 2011 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F816 (8k x 8k) / Number real images: 461 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.2 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.7 µm / Nominal magnification: 62000 |
| Sample stage | Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Reconstruction was determined by IHRSR. |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 14.4 Å Applied symmetry - Helical parameters - Δ&Phi: 116.4 ° Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.1 Å / Resolution method: OTHER / Software - Name: Spider |
| CTF correction | Details: Whole micrograph |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















