[English] 日本語
 Yorodumi
Yorodumi- EMDB-5676: Obstruction of Dengue Virus Maturation by Fab Fragments of the 2H... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5676 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Obstruction of Dengue Virus Maturation by Fab Fragments of the 2H2 Antibody | |||||||||
 Map data Map data | High-concentration 2H2 Fab complexed to immature Dengue virus back-neutralized to pH 7 after lowering to pH 6 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Dengue Maturation / Immature Dengue Antibody | |||||||||
| Biological species |   Dengue virus Dengue virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 21.0 Å | |||||||||
 Authors Authors | Wang Z / Li L / Penningtona JG / Sheng J / Yap M / Plevk P / Meng G / Sun L / Jiang W / Rossmann MG | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2013 Journal: J Virol / Year: 2013Title: Obstruction of dengue virus maturation by Fab fragments of the 2H2 antibody. Authors: Zhiqing Wang / Long Li / Janice G Pennington / Ju Sheng / Moh Lan Yap / Pavel Plevka / Geng Meng / Lei Sun / Wen Jiang / Michael G Rossmann /  Abstract: The 2H2 monoclonal antibody recognizes the precursor peptide on immature dengue virus and might therefore be a useful tool for investigating the conformational change that occurs when the immature ...The 2H2 monoclonal antibody recognizes the precursor peptide on immature dengue virus and might therefore be a useful tool for investigating the conformational change that occurs when the immature virus enters an acidic environment. During dengue virus maturation, spiky, immature, noninfectious virions change their structure to form smooth-surfaced particles in the slightly acidic environment of the trans-Golgi network, thereby allowing cellular furin to cleave the precursor-membrane proteins. The dengue virions become fully infectious when they release the cleaved precursor peptide upon reaching the neutral-pH environment of the extracellular space. Here we report on the cryo-electron microscopy structures of the immature virus complexed with the 2H2 antigen binding fragments (Fab) at different concentrations and under various pH conditions. At neutral pH and a high concentration of Fab molecules, three Fab molecules bind to three precursor-membrane proteins on each spike of the immature virus. However, at a low concentration of Fab molecules and pH 7.0, only two Fab molecules bind to each spike. Changing to a slightly acidic pH caused no detectable change of structure for the sample with a high Fab concentration but caused severe structural damage to the low-concentration sample. Therefore, the 2H2 Fab inhibits the maturation process of immature dengue virus when Fab molecules are present at a high concentration, because the three Fab molecules on each spike hold the precursor-membrane molecules together, thereby inhibiting the normal conformational change that occurs during maturation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5676.map.gz emd_5676.map.gz | 40.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5676-v30.xml emd-5676-v30.xml emd-5676.xml emd-5676.xml | 11.3 KB 11.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5676_1.jpg emd_5676_1.jpg | 190.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5676 http://ftp.pdbj.org/pub/emdb/structures/EMD-5676 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5676 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5676 | HTTPS FTP |
-Validation report
| Summary document |  emd_5676_validation.pdf.gz emd_5676_validation.pdf.gz | 78.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5676_full_validation.pdf.gz emd_5676_full_validation.pdf.gz | 77.5 KB | Display | |
| Data in XML |  emd_5676_validation.xml.gz emd_5676_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5676 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5676 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5676 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5676 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5676.map.gz / Format: CCP4 / Size: 51.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5676.map.gz / Format: CCP4 / Size: 51.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | High-concentration 2H2 Fab complexed to immature Dengue virus back-neutralized to pH 7 after lowering to pH 6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.72 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
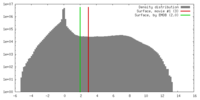
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : High-concentration Fab Fragment of 2H2 antibody binding to immatu...
| Entire | Name: High-concentration Fab Fragment of 2H2 antibody binding to immature Dengue virus back-neutralized to pH 7 after lowering to pH 6 |
|---|---|
| Components |
|
-Supramolecule #1000: High-concentration Fab Fragment of 2H2 antibody binding to immatu...
| Supramolecule | Name: High-concentration Fab Fragment of 2H2 antibody binding to immature Dengue virus back-neutralized to pH 7 after lowering to pH 6 type: sample / ID: 1000 Oligomeric state: Three Fab fragments bind to three pr on each trimeric spike Number unique components: 3 |
|---|
-Supramolecule #1: Dengue virus
| Supramolecule | Name: Dengue virus / type: virus / ID: 1 / Details: immature virus / NCBI-ID: 12637 / Sci species name: Dengue virus / Sci species strain: P16881 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Name: prM and E / Diameter: 600 Å / T number (triangulation number): 3 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 / Details: 100mM phosphate buffer |
|---|---|
| Grid | Details: Quantifoil 200 mesh |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 100 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Temperature | Min: 80 K / Max: 105 K / Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000 times magnification |
| Date | Mar 8, 2011 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON COOLSCAN / Digitization - Sampling interval: 6 µm / Number real images: 47 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 51040 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Film |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 21.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN2 / Number images used: 344 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: EMFit |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)