Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5030 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GTPase activation of elongation factor EF-Tu by the ribosome during decoding: a cryo-EM structure of the Thermus thermophilus ribosome in which the ternary complex of EF-Tu, tRNA and guanine nucleotide has been trapped on the ribosome with the antibiotic kirromycin. | |||||||||
 Map data Map data | This is an electron density map of a Thermus thermophilus ribosome complexed with mRNA, P- and E-site tRNA as well as EF-Tu.aatRNA.GDP ternary complex. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Thermus thermophilus / 70S / ribosome / EF-Tu / kirromycin / elongation factor / mRNA / Cryo / EM / GTPase / decoding | |||||||||
| Function / homology |  Function and homology information Function and homology informationtranslation elongation factor activity / regulation of translation / large ribosomal subunit / transferase activity / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / 5S rRNA binding / ribosomal large subunit assembly ...translation elongation factor activity / regulation of translation / large ribosomal subunit / transferase activity / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / 5S rRNA binding / ribosomal large subunit assembly / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / ribosome / structural constituent of ribosome / translation / ribonucleoprotein complex / GTPase activity / mRNA binding / GTP binding / RNA binding / zinc ion binding / metal ion binding / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Thermus thermophilus (bacteria) Thermus thermophilus (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 6.4 Å | |||||||||
 Authors Authors | Schuette J-C / Murphy F / Kelley AC / Weir J / Giesebrecht J / Connelll SR / Loerke J / Mielke T / Zhang W / Penczek PA ...Schuette J-C / Murphy F / Kelley AC / Weir J / Giesebrecht J / Connelll SR / Loerke J / Mielke T / Zhang W / Penczek PA / Ramakrishnan V / Spahn CMT | |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2009 Journal: EMBO J / Year: 2009Title: GTPase activation of elongation factor EF-Tu by the ribosome during decoding. Authors: Jan-Christian Schuette / Frank V Murphy / Ann C Kelley / John R Weir / Jan Giesebrecht / Sean R Connell / Justus Loerke / Thorsten Mielke / Wei Zhang / Pawel A Penczek / V Ramakrishnan / Christian M T Spahn /  Abstract: We have used single-particle reconstruction in cryo-electron microscopy to determine a structure of the Thermus thermophilus ribosome in which the ternary complex of elongation factor Tu (EF-Tu), ...We have used single-particle reconstruction in cryo-electron microscopy to determine a structure of the Thermus thermophilus ribosome in which the ternary complex of elongation factor Tu (EF-Tu), tRNA and guanine nucleotide has been trapped on the ribosome using the antibiotic kirromycin. This represents the state in the decoding process just after codon recognition by tRNA and the resulting GTP hydrolysis by EF-Tu, but before the release of EF-Tu from the ribosome. Progress in sample purification and image processing made it possible to reach a resolution of 6.4 A. Secondary structure elements in tRNA, EF-Tu and the ribosome, and even GDP and kirromycin, could all be visualized directly. The structure reveals a complex conformational rearrangement of the tRNA in the A/T state and the interactions with the functionally important switch regions of EF-Tu crucial to GTP hydrolysis. Thus, the structure provides insights into the molecular mechanism of signalling codon recognition from the decoding centre of the 30S subunit to the GTPase centre of EF-Tu. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5030.map.gz emd_5030.map.gz | 90.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5030-v30.xml emd-5030-v30.xml emd-5030.xml emd-5030.xml | 11.2 KB 11.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5030_1.tif emd_5030_1.tif | 4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5030 http://ftp.pdbj.org/pub/emdb/structures/EMD-5030 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5030 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5030 | HTTPS FTP |
-Validation report
| Summary document |  emd_5030_validation.pdf.gz emd_5030_validation.pdf.gz | 353.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5030_full_validation.pdf.gz emd_5030_full_validation.pdf.gz | 353 KB | Display | |
| Data in XML |  emd_5030_validation.xml.gz emd_5030_validation.xml.gz | 6.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5030 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5030 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5030 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5030 | HTTPS FTP |
-Related structure data
| Related structure data |  4v68MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5030.map.gz / Format: CCP4 / Size: 100.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5030.map.gz / Format: CCP4 / Size: 100.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is an electron density map of a Thermus thermophilus ribosome complexed with mRNA, P- and E-site tRNA as well as EF-Tu.aatRNA.GDP ternary complex. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.26 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
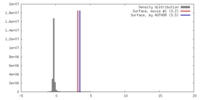
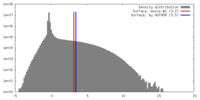
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Thermus thermophilus ribosome in which the ternary complex of elo...
| Entire | Name: Thermus thermophilus ribosome in which the ternary complex of elongation factor Tu (EF-Tu), tRNA and guanine nucleotide has been trapped on the ribosome using the antibiotic kirromycin. |
|---|---|
| Components |
|
-Supramolecule #1000: Thermus thermophilus ribosome in which the ternary complex of elo...
| Supramolecule | Name: Thermus thermophilus ribosome in which the ternary complex of elongation factor Tu (EF-Tu), tRNA and guanine nucleotide has been trapped on the ribosome using the antibiotic kirromycin. type: sample / ID: 1000 Details: The sample was purified using a His-Tag on the ternary complex, which resulted in high occupancy of the ribosome with EF-Tu ternary complex. Oligomeric state: 70S ribosome with EF-Tu ternary complex, tRNAs and mRNA Number unique components: 7 |
|---|
-Supramolecule #1: ribosome
| Supramolecule | Name: ribosome / type: complex / ID: 1 / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus (bacteria) Thermus thermophilus (bacteria) |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Staining | Type: NEGATIVE Details: Cryo-EM in vitreous ice. A VITROBOT cryo-plunger was used to prepare grids. |
|---|---|
| Grid | Details: Quantifoil grids. |
| Vitrification | Cryogen name: METHANE / Chamber humidity: 100 % / Instrument: OTHER / Details: Vitrification instrument: VITROBOT |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Details | low-dose conditions |
| Image recording | Digitization - Scanner: PRIMESCAN / Number real images: 452 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal magnification: 39000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN HELIUM |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | 586688 individual particle images were obtained from 452 micrographs. Multi-particle refinement resulted in maps with different subconformations. The deposited map represents the most prevalent conformation, based on 323688 particle images. |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 6.4 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPIDER / Number images used: 323688 |
-Atomic model buiding 1
| Initial model | PDB ID:  1vsa |
|---|---|
| Details | fitting of molecular models was performed using SITUS and SPIDER. |
| Refinement | Space: REAL |
| Output model |  PDB-4v68: |
-Atomic model buiding 2
| Initial model | PDB ID:  2hgu |
|---|---|
| Details | Protein L11. |
| Refinement | Space: REAL |
| Output model |  PDB-4v68: |
-Atomic model buiding 3
| Initial model | PDB ID:  2j00 |
|---|---|
| Refinement | Space: REAL |
| Output model |  PDB-4v68: |
-Atomic model buiding 4
| Initial model | PDB ID:  2j01 |
|---|---|
| Refinement | Space: REAL |
| Output model |  PDB-4v68: |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)