+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4538 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli DnaBC complex bound to ssDNA | |||||||||
 Map data Map data | E. coli DnaBC complex bound to ssDNA | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Helicase / helicase loader / AAA+ / RecA / REPLICATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationDnaB-DnaC complex / DnaB-DnaC-Rep-PriC complex / DnaB-DnaG complex / DnaB-DnaC-DnaT-PriA-PriC complex / DnaB-DnaC-DnaT-PriA-PriB complex / DNA helicase complex / primosome complex / DNA 5'-3' helicase / DNA replication, synthesis of primer / replisome ...DnaB-DnaC complex / DnaB-DnaC-Rep-PriC complex / DnaB-DnaG complex / DnaB-DnaC-DnaT-PriA-PriC complex / DnaB-DnaC-DnaT-PriA-PriB complex / DNA helicase complex / primosome complex / DNA 5'-3' helicase / DNA replication, synthesis of primer / replisome / DNA strand elongation involved in DNA replication / response to ionizing radiation / replication fork processing / DNA replication initiation / DNA helicase activity / helicase activity / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / 5'-3' DNA helicase activity / DNA replication / hydrolase activity / ATP hydrolysis activity / DNA binding / ATP binding / identical protein binding / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
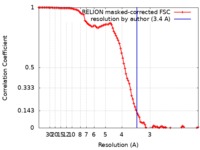
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Arias-Palomo E / Puri N | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2019 Journal: Mol Cell / Year: 2019Title: Physical Basis for the Loading of a Bacterial Replicative Helicase onto DNA. Authors: Ernesto Arias-Palomo / Neha Puri / Valerie L O'Shea Murray / Qianyun Yan / James M Berger /   Abstract: In cells, dedicated AAA+ ATPases deposit hexameric, ring-shaped helicases onto DNA to initiate chromosomal replication. To better understand the mechanisms by which helicase loading can occur, we ...In cells, dedicated AAA+ ATPases deposit hexameric, ring-shaped helicases onto DNA to initiate chromosomal replication. To better understand the mechanisms by which helicase loading can occur, we used cryo-EM to determine sub-4-Å-resolution structures of the E. coli DnaB⋅DnaC helicase⋅loader complex with nucleotide in pre- and post-DNA engagement states. In the absence of DNA, six DnaC protomers latch onto and crack open a DnaB hexamer using an extended N-terminal domain, stabilizing this conformation through nucleotide-dependent ATPase interactions. Upon binding DNA, DnaC hydrolyzes ATP, allowing DnaB to isomerize into a topologically closed, pre-translocation state competent to bind primase. Our data show how DnaC opens the DnaB ring and represses the helicase prior to DNA binding and how DnaC ATPase activity is reciprocally regulated by DnaB and DNA. Comparative analyses reveal how the helicase loading mechanism of DnaC parallels and diverges from homologous AAA+ systems involved in DNA replication and transposition. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4538.map.gz emd_4538.map.gz | 7.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4538-v30.xml emd-4538-v30.xml emd-4538.xml emd-4538.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4538_fsc.xml emd_4538_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_4538.png emd_4538.png | 75.5 KB | ||
| Filedesc metadata |  emd-4538.cif.gz emd-4538.cif.gz | 6.5 KB | ||
| Others |  emd_4538_additional.map.gz emd_4538_additional.map.gz | 70.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4538 http://ftp.pdbj.org/pub/emdb/structures/EMD-4538 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4538 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4538 | HTTPS FTP |
-Related structure data
| Related structure data |  6qemMC  4537C  6qelC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4538.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4538.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli DnaBC complex bound to ssDNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.047 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: E. coli DnaBC complex bound to ssDNA. Unsharpened map
| File | emd_4538_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli DnaBC complex bound to ssDNA. Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : E. coli DnaBC complex bound to ssDNA
| Entire | Name: E. coli DnaBC complex bound to ssDNA |
|---|---|
| Components |
|
-Supramolecule #1: E. coli DnaBC complex bound to ssDNA
| Supramolecule | Name: E. coli DnaBC complex bound to ssDNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Molecular weight | Theoretical: 490 KDa |
-Supramolecule #2: DnaBC
| Supramolecule | Name: DnaBC / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: ssDNA
| Supramolecule | Name: ssDNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Replicative DNA helicase
| Macromolecule | Name: Replicative DNA helicase / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 52.450945 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGNKPFNKQ QAEPRERDPQ VAGLKVPPHS IEAEQSVLGG LMLDNERWDD VAERVVADDF YTRPHRHIFT EMARLQESGS PIDLITLAE SLERQGQLDS VGGFAYLAEL SKNTPSAANI SAYADIVRER AVVREMISVA NEIAEAGFDP QGRTSEDLLD L AESRVFKI ...String: MAGNKPFNKQ QAEPRERDPQ VAGLKVPPHS IEAEQSVLGG LMLDNERWDD VAERVVADDF YTRPHRHIFT EMARLQESGS PIDLITLAE SLERQGQLDS VGGFAYLAEL SKNTPSAANI SAYADIVRER AVVREMISVA NEIAEAGFDP QGRTSEDLLD L AESRVFKI AESRANKDEG PKNIADVLDA TVARIEQLFQ QPHDGVTGVN TGYDDLNKKT AGLQPSDLII VAARPSMGKT TF AMNLVEN AAMLQDKPVL IFSLEMPSEQ IMMRSLASLS RVDQTKIRTG QLDDEDWARI SGTMGILLEK RNIYIDDSSG LTP TEVRSR ARRIAREHGG IGLIMIDYLQ LMRVPALSDN RTLEIAEISR SLKALAKELN VPVVALSQLN RSLEQRADKR PVNS DLRES GSIEQDADLI MFIYRDEVYH ENSDLKGIAE IIIGKQRNGP IGTVRLTFNG QWSRFDNYAG PQYDDE UniProtKB: Replicative DNA helicase DnaB |
-Macromolecule #2: DNA replication protein DnaC
| Macromolecule | Name: DNA replication protein DnaC / type: protein_or_peptide / ID: 2 / Number of copies: 6 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.973152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKNVGDLMQR LQKMMPAHIK PAFKTGEELL AWQKEQGAIR SAALERENRA MKMQRTFNRS GIRPLHQNCS FENYRVECEG QMNALSKAR QYVEEFDGNI ASFIFSGKPG TGKNHLAAAI CNELLLRGKS VLIITVADIM SAMKDTFRNS GTSEEQLLND L SNVDLLVI ...String: MKNVGDLMQR LQKMMPAHIK PAFKTGEELL AWQKEQGAIR SAALERENRA MKMQRTFNRS GIRPLHQNCS FENYRVECEG QMNALSKAR QYVEEFDGNI ASFIFSGKPG TGKNHLAAAI CNELLLRGKS VLIITVADIM SAMKDTFRNS GTSEEQLLND L SNVDLLVI DEIGVQTESK YEKVIINQIV DRRSSSKRPT GMLTNSNMEE MTKLLGERVM DRMRLGNSLW VIFNWDSYRS RV TGKEY UniProtKB: Replicative helicase loader DnaC |
-Macromolecule #3: ssDNA
| Macromolecule | Name: ssDNA / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 10.905983 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT) |
-Macromolecule #4: [[[(2R,3S,4R,5R)-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-...
| Macromolecule | Name: [[[(2R,3S,4R,5R)-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl]oxy-oxidanyl-phosphoryl]oxy-tris(fluoranyl)beryllium type: ligand / ID: 4 / Number of copies: 5 / Formula: 08T |
|---|---|
| Molecular weight | Theoretical: 492.201 Da |
| Chemical component information |  ChemComp-08T: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: AIR / Details: The grids were pre-treated with poly-lys |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-6qem: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)