+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GR89,696 bound Kappa Opioid Receptor in complex with gustducin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Kappa opioid receptor / G protein / MEMBRANE PROTEIN | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.72 Å | |||||||||
 Authors Authors | Fay JF / Che T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Ligand and G-protein selectivity in the κ-opioid receptor. Authors: Jianming Han / Jingying Zhang / Antonina L Nazarova / Sarah M Bernhard / Brian E Krumm / Lei Zhao / Jordy Homing Lam / Vipin A Rangari / Susruta Majumdar / David E Nichols / Vsevolod ...Authors: Jianming Han / Jingying Zhang / Antonina L Nazarova / Sarah M Bernhard / Brian E Krumm / Lei Zhao / Jordy Homing Lam / Vipin A Rangari / Susruta Majumdar / David E Nichols / Vsevolod Katritch / Peng Yuan / Jonathan F Fay / Tao Che /  Abstract: The κ-opioid receptor (KOR) represents a highly desirable therapeutic target for treating not only pain but also addiction and affective disorders. However, the development of KOR analgesics has ...The κ-opioid receptor (KOR) represents a highly desirable therapeutic target for treating not only pain but also addiction and affective disorders. However, the development of KOR analgesics has been hindered by the associated hallucinogenic side effects. The initiation of KOR signalling requires the G-family proteins including the conventional (G, G, G, G and G) and nonconventional (G and G) subtypes. How hallucinogens exert their actions through KOR and how KOR determines G-protein subtype selectivity are not well understood. Here we determined the active-state structures of KOR in a complex with multiple G-protein heterotrimers-G, G, G and G-using cryo-electron microscopy. The KOR-G-protein complexes are bound to hallucinogenic salvinorins or highly selective KOR agonists. Comparisons of these structures reveal molecular determinants critical for KOR-G-protein interactions as well as key elements governing G-family subtype selectivity and KOR ligand selectivity. Furthermore, the four G-protein subtypes display an intrinsically different binding affinity and allosteric activity on agonist binding at KOR. These results provide insights into the actions of opioids and G-protein-coupling specificity at KOR and establish a foundation to examine the therapeutic potential of pathway-selective agonists of KOR. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43734.map.gz emd_43734.map.gz | 85.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43734-v30.xml emd-43734-v30.xml emd-43734.xml emd-43734.xml | 13.1 KB 13.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_43734.png emd_43734.png | 29.3 KB | ||
| Filedesc metadata |  emd-43734.cif.gz emd-43734.cif.gz | 3.9 KB | ||
| Others |  emd_43734_half_map_1.map.gz emd_43734_half_map_1.map.gz emd_43734_half_map_2.map.gz emd_43734_half_map_2.map.gz | 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43734 http://ftp.pdbj.org/pub/emdb/structures/EMD-43734 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43734 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43734 | HTTPS FTP |
-Validation report
| Summary document |  emd_43734_validation.pdf.gz emd_43734_validation.pdf.gz | 828 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43734_full_validation.pdf.gz emd_43734_full_validation.pdf.gz | 827.6 KB | Display | |
| Data in XML |  emd_43734_validation.xml.gz emd_43734_validation.xml.gz | 12.9 KB | Display | |
| Data in CIF |  emd_43734_validation.cif.gz emd_43734_validation.cif.gz | 15 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43734 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43734 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43734 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43734 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43734.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43734.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
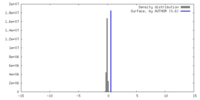
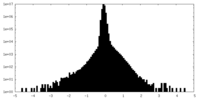
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_43734_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_43734_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GR89,696 bound Kappa Opioid Receptor in complex with gustducin
| Entire | Name: GR89,696 bound Kappa Opioid Receptor in complex with gustducin |
|---|---|
| Components |
|
-Supramolecule #1: GR89,696 bound Kappa Opioid Receptor in complex with gustducin
| Supramolecule | Name: GR89,696 bound Kappa Opioid Receptor in complex with gustducin type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 5752 / Average electron dose: 29.07 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.4 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.72 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 725000 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)