+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a bacterial gasdermin medium oval pore assembly | |||||||||
 Map data Map data | ~40mer oval pore map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | gasdermin / pore-forming protein / pyroptosis / bacteria / immunity / cell death / IMMUNE SYSTEM | |||||||||
| Function / homology | defense response to virus / plasma membrane / cytoplasm / Gasdermin bGSDM Function and homology information Function and homology information | |||||||||
| Biological species |  Vitiosangium sp. GDMCC 1.1324 (bacteria) Vitiosangium sp. GDMCC 1.1324 (bacteria) | |||||||||
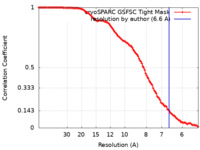
| Method | single particle reconstruction / cryo EM / Resolution: 6.6 Å | |||||||||
 Authors Authors | Johnson AG / Mayer ML / Kranzusch PJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation | Journal: bioRxiv / Year: 2023 Title: Structure and assembly of a bacterial gasdermin pore. Authors: Alex G Johnson / Megan L Mayer / Stefan L Schaefer / Nora K McNamara-Bordewick / Gerhard Hummer / Philip J Kranzusch /   Abstract: In response to pathogen infection, gasdermin (GSDM) proteins form membrane pores that induce a host cell death process called pyroptosis. Studies of human and mouse GSDM pores reveal the functions ...In response to pathogen infection, gasdermin (GSDM) proteins form membrane pores that induce a host cell death process called pyroptosis. Studies of human and mouse GSDM pores reveal the functions and architectures of 24-33 protomers assemblies, but the mechanism and evolutionary origin of membrane targeting and GSDM pore formation remain unknown. Here we determine a structure of a bacterial GSDM (bGSDM) pore and define a conserved mechanism of pore assembly. Engineering a panel of bGSDMs for site-specific proteolytic activation, we demonstrate that diverse bGSDMs form distinct pore sizes that range from smaller mammalian-like assemblies to exceptionally large pores containing >50 protomers. We determine a 3.3 Å cryo-EM structure of a bGSDM in an active slinky-like oligomeric conformation and analyze bGSDM pores in a native lipid environment to create an atomic-level model of a full 52-mer bGSDM pore. Combining our structural analysis with molecular dynamics simulations and cellular assays, our results support a stepwise model of GSDM pore assembly and suggest that a covalently bound palmitoyl can leave a hydrophobic sheath and insert into the membrane before formation of the membrane-spanning β-strand regions. These results reveal the diversity of GSDM pores found in nature and explain the function of an ancient post-translational modification in enabling programmed host cell death. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43509.map.gz emd_43509.map.gz | 75.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43509-v30.xml emd-43509-v30.xml emd-43509.xml emd-43509.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43509_fsc.xml emd_43509_fsc.xml | 11.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_43509.png emd_43509.png | 42.2 KB | ||
| Masks |  emd_43509_msk_1.map emd_43509_msk_1.map | 149.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-43509.cif.gz emd-43509.cif.gz | 5.1 KB | ||
| Others |  emd_43509_half_map_1.map.gz emd_43509_half_map_1.map.gz emd_43509_half_map_2.map.gz emd_43509_half_map_2.map.gz | 139.1 MB 139.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43509 http://ftp.pdbj.org/pub/emdb/structures/EMD-43509 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43509 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43509 | HTTPS FTP |
-Validation report
| Summary document |  emd_43509_validation.pdf.gz emd_43509_validation.pdf.gz | 810.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43509_full_validation.pdf.gz emd_43509_full_validation.pdf.gz | 810.1 KB | Display | |
| Data in XML |  emd_43509_validation.xml.gz emd_43509_validation.xml.gz | 19.7 KB | Display | |
| Data in CIF |  emd_43509_validation.cif.gz emd_43509_validation.cif.gz | 25.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43509 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43509 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43509 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43509 | HTTPS FTP |
-Related structure data
| Related structure data |  8sl0C C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43509.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43509.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ~40mer oval pore map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.66 Å | ||||||||||||||||||||||||||||||||||||
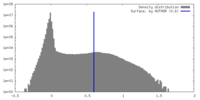
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_43509_msk_1.map emd_43509_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
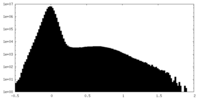
| Density Histograms |
-Half map: ~40mer oval pore half map B
| File | emd_43509_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ~40mer oval pore half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: ~40mer oval pore half map B
| File | emd_43509_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ~40mer oval pore half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Vitiosangium bGSDM in an medium oval pore assembly
| Entire | Name: Vitiosangium bGSDM in an medium oval pore assembly |
|---|---|
| Components |
|
-Supramolecule #1: Vitiosangium bGSDM in an medium oval pore assembly
| Supramolecule | Name: Vitiosangium bGSDM in an medium oval pore assembly / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Vitiosangium sp. GDMCC 1.1324 (bacteria) Vitiosangium sp. GDMCC 1.1324 (bacteria) |
-Macromolecule #1: Vitiosangium bacterial gasdermin
| Macromolecule | Name: Vitiosangium bacterial gasdermin / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Vitiosangium sp. GDMCC 1.1324 (bacteria) Vitiosangium sp. GDMCC 1.1324 (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: SGLCSDPAIT YLKRLGYNVV RLPREGIQPL HLLGQQRGTV EYLGSLEKLI TQPPSEPPAI TRDQAAAGIN GQKTENLSFS IGINILKSVL AQFGAGAGIE AQYNQARKVR FEFSNVLADS VEPLAVGQFL KMAEVDADNP VLKQYVLGNG RLYVITQVIK SNEFTVAAEK ...String: SGLCSDPAIT YLKRLGYNVV RLPREGIQPL HLLGQQRGTV EYLGSLEKLI TQPPSEPPAI TRDQAAAGIN GQKTENLSFS IGINILKSVL AQFGAGAGIE AQYNQARKVR FEFSNVLADS VEPLAVGQFL KMAEVDADNP VLKQYVLGNG RLYVITQVIK SNEFTVAAEK SGGGSIQLDV PEIQKVVGGK LKVEASVSSQ STVTYKGEKQ LVFGFKCFEI GVKNGEITLF ASQLVPR UniProtKB: Gasdermin bGSDM |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 150 mM NaCl, 20 mM HEPES-HOH (pH 7.5), 50 mM HECAMEG | ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 2 | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | The sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 33411 / Average electron dose: 53.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)