+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | A DARPin displayed on a designed tetrahedral protein scaffold | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | scaffold / tetrahedral / darpin / symmetrical / DE NOVO PROTEIN | |||||||||
| Biological species | synthetic construct (others) | |||||||||
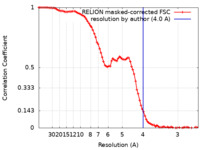
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Suder DS / Gonen S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Int J Mol Sci / Year: 2024 Journal: Int J Mol Sci / Year: 2024Title: Mitigating the Blurring Effect of CryoEM Averaging on a Flexible and Highly Symmetric Protein Complex through Sub-Particle Reconstruction. Authors: Diana S Suder / Shane Gonen /  Abstract: Many macromolecules are inherently flexible as a feature of their structure and function. During single-particle CryoEM processing, flexible protein regions can be detrimental to high-resolution ...Many macromolecules are inherently flexible as a feature of their structure and function. During single-particle CryoEM processing, flexible protein regions can be detrimental to high-resolution reconstruction as signals from thousands of particles are averaged together. This "blurring" effect can be difficult to overcome and is possibly more pronounced when averaging highly symmetric complexes. Approaches to mitigating flexibility during CryoEM processing are becoming increasingly critical as the technique advances and is applied to more dynamic proteins and complexes. Here, we detail the use of sub-particle averaging and signal subtraction techniques to precisely target and resolve flexible DARPin protein attachments on a designed tetrahedrally symmetric protein scaffold called DARP14. Particles are first aligned as full complexes, and then the symmetry is reduced by alignment and focused refinement of the constituent subunits. The final reconstructions we obtained were vastly improved over the fully symmetric reconstructions, with observable secondary structure and side-chain placement. Additionally, we were also able to reconstruct the core region of the scaffold to 2.7 Å. The data processing protocol outlined here is applicable to other dynamic and symmetric protein complexes, and our improved maps could allow for new structure-guided variant designs of DARP14. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43167.map.gz emd_43167.map.gz | 48.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43167-v30.xml emd-43167-v30.xml emd-43167.xml emd-43167.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43167_fsc.xml emd_43167_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_43167.png emd_43167.png | 69.5 KB | ||
| Filedesc metadata |  emd-43167.cif.gz emd-43167.cif.gz | 5.7 KB | ||
| Others |  emd_43167_half_map_1.map.gz emd_43167_half_map_1.map.gz emd_43167_half_map_2.map.gz emd_43167_half_map_2.map.gz | 40.7 MB 40.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43167 http://ftp.pdbj.org/pub/emdb/structures/EMD-43167 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43167 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43167 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43167.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43167.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_43167_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_43167_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : DARPin with a helical connection to a chain of a designed protein...
| Entire | Name: DARPin with a helical connection to a chain of a designed protein scaffold |
|---|---|
| Components |
|
-Supramolecule #1: DARPin with a helical connection to a chain of a designed protein...
| Supramolecule | Name: DARPin with a helical connection to a chain of a designed protein scaffold type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: DARPin protein scaffold
| Macromolecule | Name: DARPin protein scaffold / type: protein_or_peptide / ID: 1 Details: with a helical connection to a chain of a designed protein scaffold Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 32.208723 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KDSPIIEANG TLDELTSFIG EAKHYVDEEM KGILEEIQND IYKIMGEIGS KGKIEGISEE RIAWLLKLIL RYMEMVNLKS FVLPGGTLE SAKLDVCRTI ARRALRKVLT VTREFGIGAE AAAYLLALSD LLFLLARVIE IELGKKLLEA ARAGQDDEVR I LMANGADV ...String: KDSPIIEANG TLDELTSFIG EAKHYVDEEM KGILEEIQND IYKIMGEIGS KGKIEGISEE RIAWLLKLIL RYMEMVNLKS FVLPGGTLE SAKLDVCRTI ARRALRKVLT VTREFGIGAE AAAYLLALSD LLFLLARVIE IELGKKLLEA ARAGQDDEVR I LMANGADV NAHDDQGSTP LHLAAWIGHP EIVEVLLKHG ADVNARDTDG WTPLHLAADN GHLEIVEVLL KYGADVNAQD AY GLTPLHL AADRGHLEIV EVLLKHGADV NAQDKFGKTA FDISIDNGNE DLAEILQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)