[English] 日本語
 Yorodumi
Yorodumi- EMDB-42442: CryoEM Structure of Allosterically Switchable De Novo Protein sr3... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Structure of Allosterically Switchable De Novo Protein sr322, In Closed State without Effector Peptide | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Allosterically Switchable Protein / sr322 / closed state / DE NOVO PROTEIN | |||||||||
| Biological species | unidentified (others) | |||||||||
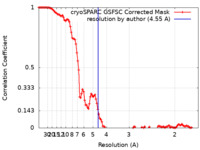
| Method | single particle reconstruction / cryo EM / Resolution: 4.55 Å | |||||||||
 Authors Authors | Weidle C / Borst A | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: De novo design of allosterically switchable protein assemblies. Authors: Arvind Pillai / Abbas Idris / Annika Philomin / Connor Weidle / Rebecca Skotheim / Philip J Y Leung / Adam Broerman / Cullen Demakis / Andrew J Borst / Florian Praetorius / David Baker /   Abstract: Allosteric modulation of protein function, wherein the binding of an effector to a protein triggers conformational changes at distant functional sites, plays a central part in the control of ...Allosteric modulation of protein function, wherein the binding of an effector to a protein triggers conformational changes at distant functional sites, plays a central part in the control of metabolism and cell signalling. There has been considerable interest in designing allosteric systems, both to gain insight into the mechanisms underlying such 'action at a distance' modulation and to create synthetic proteins whose functions can be regulated by effectors. However, emulating the subtle conformational changes distributed across many residues, characteristic of natural allosteric proteins, is a significant challenge. Here, inspired by the classic Monod-Wyman-Changeux model of cooperativity, we investigate the de novo design of allostery through rigid-body coupling of peptide-switchable hinge modules to protein interfaces that direct the formation of alternative oligomeric states. We find that this approach can be used to generate a wide variety of allosterically switchable systems, including cyclic rings that incorporate or eject subunits in response to peptide binding and dihedral cages that undergo effector-induced disassembly. Size-exclusion chromatography, mass photometry and electron microscopy reveal that these designed allosteric protein assemblies closely resemble the design models in both the presence and absence of peptide effectors and can have ligand-binding cooperativity comparable to classic natural systems such as haemoglobin. Our results indicate that allostery can arise from global coupling of the energetics of protein substructures without optimized side-chain-side-chain allosteric communication pathways and provide a roadmap for generating allosterically triggerable delivery systems, protein nanomachines and cellular feedback control circuitry. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42442.map.gz emd_42442.map.gz | 134 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42442-v30.xml emd-42442-v30.xml emd-42442.xml emd-42442.xml | 23.8 KB 23.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42442_fsc.xml emd_42442_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_42442.png emd_42442.png | 162.2 KB | ||
| Filedesc metadata |  emd-42442.cif.gz emd-42442.cif.gz | 7 KB | ||
| Others |  emd_42442_additional_1.map.gz emd_42442_additional_1.map.gz emd_42442_half_map_1.map.gz emd_42442_half_map_1.map.gz emd_42442_half_map_2.map.gz emd_42442_half_map_2.map.gz | 140.5 MB 138.7 MB 138.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42442 http://ftp.pdbj.org/pub/emdb/structures/EMD-42442 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42442 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42442 | HTTPS FTP |
-Related structure data
| Related structure data |  8up1MC  8ureC  8utmC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42442.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42442.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: C1 map at 6.54 Angstrom
| File | emd_42442_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 map at 6.54 Angstrom | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_42442_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_42442_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : sr322
| Entire | Name: sr322 |
|---|---|
| Components |
|
-Supramolecule #1: sr322
| Supramolecule | Name: sr322 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Complex is made up of 4 monomeric proteins, and is purified as a 4 sided multimer. |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 163.296 MDa |
-Macromolecule #1: De Novo Protein sr322
| Macromolecule | Name: De Novo Protein sr322 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 40.089242 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SGTVTFDITN ISHKAIDIIL KVVLGIAEHE GTEVTFHSER GQLQIEVKNL HEEDKRLIEQ AIEAARLADS PDPESVARAV ELLTKVAKA STNTELIQFI VKELLELARK LTDPKDLAKV LDSISELLTE LALKTGDPTA ALAAMVAHIA ELVVRLALMA E RTHPGSEI ...String: SGTVTFDITN ISHKAIDIIL KVVLGIAEHE GTEVTFHSER GQLQIEVKNL HEEDKRLIEQ AIEAARLADS PDPESVARAV ELLTKVAKA STNTELIQFI VKELLELARK LTDPKDLAKV LDSISELLTE LALKTGDPTA ALAAMVAHIA ELVVRLALMA E RTHPGSEI VKKAVKLVQE VAEEVLEAAQ LMLEKPNSDE VAKKLEEVAK KAIEACIELQ QILEAWAKER GDQDLLREVR EH KLQILTI AVAYKAAQMG VTVLKHTHGW VVFLVILGLH KQQAEQLLRF VHRVAHALGV TLSITFSGDI VVIAVTVGAS EEE KKEVRK IVKEIAKQLR HAETEEEAKE IVQRVIEEWQ EEGGSG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.971 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: 150 mM NaCl, 40 mM Tris pH 8.0 | |||||||||
| Grid | Model: C-flat / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 40 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 39.0 kPa / Details: -15mA | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 4213 / Average exposure time: 5.0 sec. / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Initial model type: in silico model |
|---|---|
| Details | Initial fit was done using Chimera using Rigid body from In silico model. Then Isolde and Namdinator were used. Coot and Phenix were also used. |
| Refinement | Protocol: FLEXIBLE FIT |
| Output model |  PDB-8up1: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)