+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | VMAT1 dimer in unbound form and with reserpine | |||||||||
 マップデータ マップデータ | VMAT1 dimer in unbound form and with reserpine | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | VMAT / SLC18 / vascular monoamine transporter / monoamines / neurotransmitters / reserpine / MEMBRANE PROTEIN | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.5 Å | |||||||||
 データ登録者 データ登録者 | Ye J / Liu B / Li W | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2024 ジャーナル: Nature / 年: 2024タイトル: Structural insights into vesicular monoamine storage and drug interactions. 著者: Jin Ye / Huaping Chen / Kaituo Wang / Yi Wang / Aaron Ammerman / Samjhana Awasthi / Jinbin Xu / Bin Liu / Weikai Li /   要旨: Biogenic monoamines-vital transmitters orchestrating neurological, endocrinal and immunological functions-are stored in secretory vesicles by vesicular monoamine transporters (VMATs) for controlled ...Biogenic monoamines-vital transmitters orchestrating neurological, endocrinal and immunological functions-are stored in secretory vesicles by vesicular monoamine transporters (VMATs) for controlled quantal release. Harnessing proton antiport, VMATs enrich monoamines around 10,000-fold and sequester neurotoxicants to protect neurons. VMATs are targeted by an arsenal of therapeutic drugs and imaging agents to treat and monitor neurodegenerative disorders, hypertension and drug addiction. However, the structural mechanisms underlying these actions remain unclear. Here we report eight cryo-electron microscopy structures of human VMAT1 in unbound form and in complex with four monoamines (dopamine, noradrenaline, serotonin and histamine), the Parkinsonism-inducing MPP, the psychostimulant amphetamine and the antihypertensive drug reserpine. Reserpine binding captures a cytoplasmic-open conformation, whereas the other structures show a lumenal-open conformation stabilized by extensive gating interactions. The favoured transition to this lumenal-open state contributes to monoamine accumulation, while protonation facilitates the cytoplasmic-open transition and concurrently prevents monoamine binding to avoid unintended depletion. Monoamines and neurotoxicants share a binding pocket that possesses polar sites for specificity and a wrist-and-fist shape for versatility. Variations in this pocket explain substrate preferences across the SLC18 family. Overall, these structural insights and supporting functional studies elucidate the mechanism of vesicular monoamine transport and provide the basis to develop therapeutics for neurodegenerative diseases and substance abuse. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_41238.map.gz emd_41238.map.gz | 110.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-41238-v30.xml emd-41238-v30.xml emd-41238.xml emd-41238.xml | 15.6 KB 15.6 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
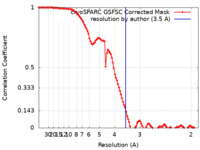
| FSC (解像度算出) |  emd_41238_fsc.xml emd_41238_fsc.xml | 10.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_41238.png emd_41238.png | 123.9 KB | ||
| Filedesc metadata |  emd-41238.cif.gz emd-41238.cif.gz | 5.8 KB | ||
| その他 |  emd_41238_half_map_1.map.gz emd_41238_half_map_1.map.gz emd_41238_half_map_2.map.gz emd_41238_half_map_2.map.gz | 116.1 MB 116.1 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41238 http://ftp.pdbj.org/pub/emdb/structures/EMD-41238 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41238 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41238 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_41238.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_41238.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | VMAT1 dimer in unbound form and with reserpine | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.94 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: VMAT1 dimer in unbound form and with reserpine
| ファイル | emd_41238_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | VMAT1 dimer in unbound form and with reserpine | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: VMAT1 dimer in unbound form and with reserpine
| ファイル | emd_41238_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | VMAT1 dimer in unbound form and with reserpine | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Complex of VMAT1 dimer in unbound form and with reserpine
| 全体 | 名称: Complex of VMAT1 dimer in unbound form and with reserpine |
|---|---|
| 要素 |
|
-超分子 #1: Complex of VMAT1 dimer in unbound form and with reserpine
| 超分子 | 名称: Complex of VMAT1 dimer in unbound form and with reserpine タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 105.2 KDa |
-分子 #1: VMAT1 dimer in unbound form and with reserpine
| 分子 | 名称: VMAT1 dimer in unbound form and with reserpine / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 50.565641 KDa |
| 組換発現 | 生物種:  Komagataella pastoris (菌類) Komagataella pastoris (菌類) |
| 配列 | 文字列: MLRTILDAPQ RLLKEGRASR QLVLVVVFVA LLLDNMLFTV VVPIVPTFLY DMEFKEVNSS LHLGHAGSNC LQGTGFLEEE ITRVGVLFA SKAVMQLLVN PFVGPLTNRI GYHIPMFAGF VIMFLSTVMF AFSGTYTLLF VARTLQGIGS SFSSVAGLGM L ASVYTDDH ...文字列: MLRTILDAPQ RLLKEGRASR QLVLVVVFVA LLLDNMLFTV VVPIVPTFLY DMEFKEVNSS LHLGHAGSNC LQGTGFLEEE ITRVGVLFA SKAVMQLLVN PFVGPLTNRI GYHIPMFAGF VIMFLSTVMF AFSGTYTLLF VARTLQGIGS SFSSVAGLGM L ASVYTDDH ERGRAMGTAL GGLALGLLVG APFGSVMYEF VGKSAPFLIL AFLALLDGAL QLCILQPSKV SPESAKGTPL FM LLKDPYI LVAAGSICFA NMGVAILEPT LPIWMMQTMC SPKWQLGLAF LPASVSYLIG TNLFGVLANK MGRWLCSLIG MLV VGTSLL CVPLAHNIFG LIGPNAGLGL AIGMVDSSMM PIMGHLVDLR HTSVYGSVYA IADVAFCMGF AIGPSTGGAI VKAI GFPWL MVITGVINIV YAPLCYYLRS PPAKEEKLAI LSQDCPMETR MYATQKPTKE FPLGEDSDEE PDHEE |
-分子 #2: reserpine
| 分子 | 名称: reserpine / タイプ: ligand / ID: 2 / コピー数: 1 / 式: YHR |
|---|---|
| 分子量 | 理論値: 608.679 Da |
| Chemical component information |  ChemComp-YHR: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 8 |
|---|---|
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: COPPER / メッシュ: 300 / 前処理 - タイプ: GLOW DISCHARGE |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS GLACIOS |
|---|---|
| 温度 | 最低: 63.0 K / 最高: 77.2 K |
| 撮影 | フィルム・検出器のモデル: FEI FALCON IV (4k x 4k) 撮影したグリッド数: 1 / 実像数: 4301 / 平均露光時間: 9.36 sec. / 平均電子線量: 47.88 e/Å2 |
| 電子線 | 加速電圧: 200 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 50.0 µm / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 2.2 µm / 最小 デフォーカス(公称値): 1.0 µm / 倍率(公称値): 150000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
 ムービー
ムービー コントローラー
コントローラー


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)