[English] 日本語
 Yorodumi
Yorodumi- EMDB-40905: Cryo-EM Structure of NINJ1 Filament at 2.75 Angstrom Resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of NINJ1 Filament at 2.75 Angstrom Resolution | |||||||||
 Map data Map data | Cryo-EM Map of NINJ1 Filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NINJ1 Filament / Plasma Membrane Rupture Protein / Cholesterol Binding Protein / Lipid Binding Protein / Membrane Protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationcell adhesion mediator activity / leukocyte chemotaxis involved in inflammatory response / membrane destabilizing activity / positive regulation of toll-like receptor 4 signaling pathway / tissue regeneration / muscle cell differentiation / programmed cell death / heterotypic cell-cell adhesion / pyroptotic inflammatory response / synaptic membrane ...cell adhesion mediator activity / leukocyte chemotaxis involved in inflammatory response / membrane destabilizing activity / positive regulation of toll-like receptor 4 signaling pathway / tissue regeneration / muscle cell differentiation / programmed cell death / heterotypic cell-cell adhesion / pyroptotic inflammatory response / synaptic membrane / lipopolysaccharide binding / protein homooligomerization / positive regulation of inflammatory response / positive regulation of angiogenesis / nervous system development / angiogenesis / killing of cells of another organism / cell adhesion / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
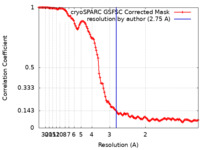
| Method | single particle reconstruction / cryo EM / Resolution: 2.75 Å | |||||||||
 Authors Authors | Sahoo B / Dai X | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Biorxiv / Year: 2023 Journal: Biorxiv / Year: 2023Title: How NINJ1 mediates plasma membrane rupture and why NINJ2 cannot Authors: Sahoo B / Mou Z / Liu W / Dubyak G / Dai X | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40905.map.gz emd_40905.map.gz | 168 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40905-v30.xml emd-40905-v30.xml emd-40905.xml emd-40905.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40905_fsc.xml emd_40905_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_40905.png emd_40905.png | 96.7 KB | ||
| Filedesc metadata |  emd-40905.cif.gz emd-40905.cif.gz | 5.1 KB | ||
| Others |  emd_40905_half_map_1.map.gz emd_40905_half_map_1.map.gz emd_40905_half_map_2.map.gz emd_40905_half_map_2.map.gz | 165.4 MB 165.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40905 http://ftp.pdbj.org/pub/emdb/structures/EMD-40905 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40905 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40905 | HTTPS FTP |
-Validation report
| Summary document |  emd_40905_validation.pdf.gz emd_40905_validation.pdf.gz | 950.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40905_full_validation.pdf.gz emd_40905_full_validation.pdf.gz | 950.4 KB | Display | |
| Data in XML |  emd_40905_validation.xml.gz emd_40905_validation.xml.gz | 20.5 KB | Display | |
| Data in CIF |  emd_40905_validation.cif.gz emd_40905_validation.cif.gz | 26.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40905 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40905 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40905 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40905 | HTTPS FTP |
-Related structure data
| Related structure data |  8szaMC  8szbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40905.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40905.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM Map of NINJ1 Filament | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.66 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
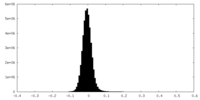
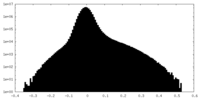
-Half map: Cryo-EM Half Map A of NINJ1 Filament
| File | emd_40905_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM Half Map A of NINJ1 Filament | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
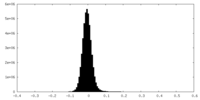
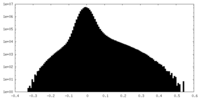
-Half map: Cryo-EM Half Map B of NINJ1 Filament
| File | emd_40905_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM Half Map B of NINJ1 Filament | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ninjurin-1 in complex with Cholesterol
| Entire | Name: Ninjurin-1 in complex with Cholesterol |
|---|---|
| Components |
|
-Supramolecule #1: Ninjurin-1 in complex with Cholesterol
| Supramolecule | Name: Ninjurin-1 in complex with Cholesterol / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Ninjurin-1
| Macromolecule | Name: Ninjurin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 19.276896 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDHDGDY KDHDIDYKDD DDKGSGDSGT EEYELNGGLP PGTPGSPDAS PARWGWRHGP INVNHYASKK SAAESMLDIA LLMANASQL KAVVEQGPSF AFYVPLVVLI SISLVLQIGV GVLLIFLVKY DLNNPAKHAK LDFLNNLATG LVFIIVVVNI F ITAFGVQK PLMDMAPQQ UniProtKB: Ninjurin-1 |
-Macromolecule #2: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 2 / Number of copies: 6 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z
Z Y
Y X
X