[English] 日本語
 Yorodumi
Yorodumi- EMDB-39767: Cryo-EM structure of CTR-bound type 7 CRISPR-Cas complex at subst... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of CTR-bound type 7 CRISPR-Cas complex at substrate-engaged state 1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | a protein complex / ANTIVIRAL PROTEIN | |||||||||
| Biological species | metagenome (others) | |||||||||
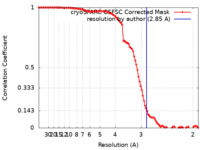
| Method | single particle reconstruction / cryo EM / Resolution: 2.85 Å | |||||||||
 Authors Authors | Zhang H / Deng Z / Li X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of CTR-bound type 7 CRISPR-Cas complex at substrate-engaged state 1 Authors: Li X / Deng Z / Zhang H | |||||||||
| History |
|
- Structure visualization
Structure visualization
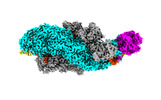
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39767.map.gz emd_39767.map.gz | 97.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39767-v30.xml emd-39767-v30.xml emd-39767.xml emd-39767.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_39767_fsc.xml emd_39767_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_39767.png emd_39767.png | 65.4 KB | ||
| Filedesc metadata |  emd-39767.cif.gz emd-39767.cif.gz | 5.9 KB | ||
| Others |  emd_39767_half_map_1.map.gz emd_39767_half_map_1.map.gz emd_39767_half_map_2.map.gz emd_39767_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39767 http://ftp.pdbj.org/pub/emdb/structures/EMD-39767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39767 | HTTPS FTP |
-Related structure data
| Related structure data |  8z4l M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_39767.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39767.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.95 Å | ||||||||||||||||||||||||||||||||||||
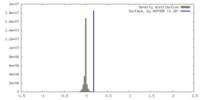
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_39767_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
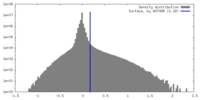
| Density Histograms |
-Half map: #1
| File | emd_39767_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
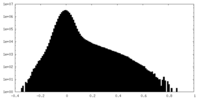
| Density Histograms |
- Sample components
Sample components
-Entire : a protein
| Entire | Name: a protein |
|---|---|
| Components |
|
-Supramolecule #1: a protein
| Supramolecule | Name: a protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism: metagenome (others) |
-Macromolecule #1: a protein
| Macromolecule | Name: a protein / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 22.255604 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAKTMKKIYV TMKTLSPLYT GEVRREDKEA AQKRVNFPVR KTATNKVLIP FKGALRSALE IMLKAKGENV CDTGESRARP CGRCVTCSL FGSMGRAGRA SVDFLISNDT KEQIVRESTH LRIERQTKSA SDTFKGEEVI EGATFTATIT ISNPQEKDLS L IQSALKFI ...String: MAKTMKKIYV TMKTLSPLYT GEVRREDKEA AQKRVNFPVR KTATNKVLIP FKGALRSALE IMLKAKGENV CDTGESRARP CGRCVTCSL FGSMGRAGRA SVDFLISNDT KEQIVRESTH LRIERQTKSA SDTFKGEEVI EGATFTATIT ISNPQEKDLS L IQSALKFI EENGIGGWLN KGYGRVSFEV KSEDVATDRF LK |
-Macromolecule #2: a protein
| Macromolecule | Name: a protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 27.139533 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKEIKGILES ITGFSIPLDN GEYALYPAGR HLRGAIGYIA FNLDLPISSK FLDFDFDDII FRDLLPISKC GKIFYPEKNS NSLKCPSCN EIYGSSVLRN IMARGLSYKE VIEGKKYRLS IIVKDEKYLN EMEAIIRYIL SYGIYLGNKV SKGYGKFKIK E YSIVDILP ...String: MKEIKGILES ITGFSIPLDN GEYALYPAGR HLRGAIGYIA FNLDLPISSK FLDFDFDDII FRDLLPISKC GKIFYPEKNS NSLKCPSCN EIYGSSVLRN IMARGLSYKE VIEGKKYRLS IIVKDEKYLN EMEAIIRYIL SYGIYLGNKV SKGYGKFKIK E YSIVDILP VKDSEVLLLS DAIIDNGEKD IVFSKKEISS SKFEIIRKRG KAKGDIIRDN NHNGFYIGKY GGLGFGEIIS LK |
-Macromolecule #3: a protein
| Macromolecule | Name: a protein / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 70.468898 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIKFIGGASK VTGSAFLLET GNAKILIDCG IEQEKGIEKD NNEIIEKKIN EIGKADICIL THAHLAASGL VPLLVKKRKV NKIISTPAT KELCRLLFND FQRIQEENND IPLYSYDDIE SSFEIWDEID DRNTIELFDT KITFYNNSHI IGSVSVFIET H NGNYLFSG ...String: MIKFIGGASK VTGSAFLLET GNAKILIDCG IEQEKGIEKD NNEIIEKKIN EIGKADICIL THAHLAASGL VPLLVKKRKV NKIISTPAT KELCRLLFND FQRIQEENND IPLYSYDDIE SSFEIWDEID DRNTIELFDT KITFYNNSHI IGSVSVFIET H NGNYLFSG DIGSKLQQLM DYPPDMPDGN VDYLILESTY GNKSHDSSDR DRLLEIAKTT CENGGKVLIP SFAIGRLQEV LY TFSNYNF NFPVYIDSPM GSKVTNLIKE YNIYLKKKLR RLSITDDLFN NKYIAINTSN QSKELSNSKE PAVIISASGM LEG GRILNH LEQIKNDENS TLIFVGYQAQ NTRGRKILDG EEKVRCRIEK LNSFSAHADQ DELIDYIERL KYTPYKVFLV HGEK EQREI LAKRIISKKI RVELPENYSQ GKEILIEKKV VLNINTDNMC NFASYRLMPF SGFIVEKDDR IEINDKNWFD MIWNE EYNK MRSQIVAEDF STDQNEDSMA LPDMSHDKII ENIEYLFNIK ILSKNRIKEF WEEFCKGQKA AIKYITQVHR KNPNTG RRN WNPPEGDFTD NEIEKLYETA YNTLLSLIKY DKNKVYNILI NFNPKL |
-Macromolecule #4: RNA (49-MER)
| Macromolecule | Name: RNA (49-MER) / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 19.142277 KDa |
| Sequence | String: GGUUAAAACU CUUCUCAUGC UGGAUUCGAA AUUAGGUGCG CUUCGCGUUU AAGUCCCAUA |
-Macromolecule #5: RNA (40-MER)
| Macromolecule | Name: RNA (40-MER) / type: rna / ID: 5 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 19.354707 KDa |
| Sequence | String: GAACAGAAGA ACACCUAAAC GCGAAGCGCA CCUAAUUUCG AAUCCAGCAU GAGAAGCUAA |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 9 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)